|
|
1.IntroductionIn an effort to better visualize biological tissue structure, polarized light imaging has been used extensively to enhance spatial resolution and to provide contrast that cannot be achieved using ordinary light alone.1–7 In addition, polarized light-based techniques (polarimetry) have allowed tissue characterization via intrinsic properties such as birefringence (arising, for example, from fibrous connective tissue networks containing collagen and elastin), orientation (arising from structural organization), and optical activity (arising from the presence of chiral molecules such as glucose).8–10 Polarimetry-based metrics have demonstrated sensitivity to changes in tissue structure and composition and may be closely associated with underlying pathology. For instance, scar formation in the myocardium has been linked to a decrease in the birefringence signal.8–10 One of the limitations to polarized light-based techniques in optically thick turbid media (such as bulk tissue) is the effect of multiple scattering, which effectively randomizes the photon direction, phase coherence, and polarization. Consequently, only a fraction of the light emerging from a sample in particular detection geometry will remain polarized. Since polarimetry relies on measuring, quantifying, and interpreting the surviving polarized light fraction, examining the depolarization rates in different tissue types and relating these to tissue structure and pathology are of great importance. Previously, several studies have concentrated on light depolarization in biological tissues. Demos et al. have suggested using the degree of polarization for discriminating between different types of tissues.11 Jacques et al. have used polarized light imaging to distinguish superficial backscattered light from the collagen fibers versus highly depolarized light from deep skin layers.12 The same group also has compared chicken liver and breast and porcine muscle and skin with a transmission mode polarimetry system, and it reported that linearly polarized light depolarizes faster in (anisotropic) tissues with birefringence, compared to isotropic tissues.13 Vitkin et al. have demonstrated in vivo that light that is diffusively scattered by human skin retains significant polarization in exact backscattering directions, and the degree of polarization increases with increased absorption.14 Sankaran et al. have investigated the degree of polarization in five porcine tissues (fat, tendon, blood, myocardium muscle, and artery) of different thicknesses (0.1 to 2 mm) in transmission geometry. They have shown that the Rayleigh-scattering regime is dominant for all the tissues except blood, with the degree of linear polarization exceeding the degree of circular polarization.15 More recently, Antonelli et al. compared the Mueller matrix in the backscattering direction from healthy and cancerous human colon biopsy samples, indicating that the cancerous samples were less depolarizing.16 A common theme in these studies is to quantify and understand the depolarization mechanisms in tissues and to use depolarization as a diagnostic tool. The current article thus describes our efforts in measuring, quantifying, and understanding depolarization behavior in a variety of bulk porcine tissues in the context of their measured optical properties. A commonly used approach to characterize tissues is by measuring their optical properties, specifically the reduced scattering coefficient and the absorption coefficient . The reduced scattering coefficient is the probability of the photons scattering in the forward direction per infinitesimal path-length. The absorption coefficient is the probability of the photons absorption per infinitesimal path-length. Several suitable methods have been developed, with their relative advantages, disadvantages, and ranges of validity closely examined.17 In the current study, we performed polarimetric and optical property measurements and analysis on optically thick samples from six different porcine tissue types. The isotropic (low birefringence) tissues examined were the liver, kidney cortex, and brain. The anisotropic (high birefringence) tissues were the myocardium muscle, loin muscle, and tendon. To quantify the dependence of the depolarization rates on source-sample-detector geometry, we performed polarimetric imaging in both transmission and backscattering modes. The paper is organized as follows: in “Materials and Methods” section, we outline sample preparation, the polarimetric imaging setup, and the methodology for measuring tissue optical properties; we also review Mueller matrix derivation, calculations, and decomposition. In “Results and Discussion,” we present the depolarization findings of the six different tissue types, correlate these with their measured optical properties, and discuss the meaning and implications of these correlative trends. Conclusions and future directions are presented in the final section. 2.Materials and MethodsWe obtained fresh porcine tissues following sacrifice from a local abattoir. All the samples were imaged and characterized within three hours after sacrifice. Samples were moistened periodically with phosphate-buffered saline solution (0.02 M) to maintain freshness; this avoids potential complications associated with formalin fixation and other tissue handling or preparation methods.18 Three previously reported low-birefringence () tissues (liver, kidney cortex, and brain) and three highly birefringent (anisotropic) tissues (myocardium muscle, loin muscle, and tendon) were chosen. Liver was selected as an isotropic and highly absorbent tissue.17,19 In the case of the kidney, we examined the cortex, which is known to be optically isotropic.17,20 Brain was chosen as a highly scattering, low-absorbing medium considered to be non-birefringent.17,21 From the heart, we chose the myocardium, which is a highly birefringent tissue composed of oriented cardiac muscle cells.8,17 Loin muscle was another anisotropic tissue examined, where the myosin fibers are known to be loosely aligned.17,22 Tendon was chosen because it exhibits the same range of the optical properties as the brain but is anisotropic, stemming from parallel arrays of closely packed collagen.15 To measure tissue optical properties of the bulk tissues, a fiberoptic reflectance probe with multiple source-collector separations was used. The optical properties and were extracted using the established technique of spatially resolved, steady-state diffuse reflectance (based on diffusion theory) as outlined by Farrell et al.23,24 Briefly, a multifiber probe with source-collector distances, , of 0.8, 1.5, 2.4, 4.0, and 5.3 mm was connected to an optical control system that consisted of a diode laser (Thorlabs, Newton, NJ) at 635 nm, an optical multiplexer (MPM-2000, Ocean Optics, Dunedin, FL), and a visible-light spectrometer (S2000, Ocean Optics). The collector fiber was connected to the spectrometer. The multiplexer was used to index the laser beam from fiber-to-fiber so as to achieve the various source-collector distances required for the spatially resolved diffuse reflectance algorithm. The well-known diffuse reflectance equations were employed to model the reflectance at each .23,24 The probe was calibrated against a diluted Intralipid (Fresenius Kabi, Sweden) solution with known optical properties and validated against a series of Intralipid-dye phantoms with and . The two optical property coefficients, and , were computed from the five reflectance measurements by solving an inverse problem. It is worth noting that the algorithm did not utilize some of the reflectance measurements in the case of low because the diffusion theory reflectance equation breaks down at low for low , as outlined in Kim et al.25 Hence, the algorithm would omit the reflectance value at, say, for , and use only four reflectance measurements to solve in the inverse problem in this scattering range. With this probe, we measured the optical properties of the six different tissues, periodically examining a well-characterized scattering solution of Intralipid as a calibration/system validation control phantom. For polarimetric measurements, two slabs of each type of tissue, one with 2 mm of thickness and one with 1 cm of thickness, were sectioned. No formalin fixation was used to avoid possible crosslinking-induced polarization property changes,18 so the results apply to freshly excised tissues. Cutting the fresh tissue with a blade to a uniform thickness is challenging; therefore, several sections were made and the three most uniform slabs with the desired thicknesses were chosen for the experiment. The resultant tissue slabs were placed between two glass slides. The 2-mm samples were imaged in both transmission and backscattering geometry, while the 1-cm samples were imaged only in backscattering mode (not surprisingly, complete polarization loss was observed in transmission for all 1-cm tissues). The polarimetric imaging setup shown in Fig. 1 was used. A diode laser (Thorlabs) with a wavelength of 635 nm was used to illuminate the sample. The polarizer, P1, and the removable quarter waveplate, WP1, were used to generate different linear and circular input polarization states. The first lens, L1, was used to project a 1-cm-diameter spot size onto the sample. Light exiting the sample in a particular detection direction then was channelled through an output arm consisting of a polarizer, P2, a removable waveplate, WP2, and a collecting lens, L2, that focused the light onto a charge-coupled device (CCD) camera (CoolSNAPK4, Photometrics, Tuscon, AZ). The output polarization was selected by P2 and WP2. Imaging was performed in two different geometries. In transmission mode, the output arm was placed collinear to the incident beam. In backscattering mode, light was collected by the output arm at an angle of 155 deg off-axis relative to the incident beam (25 deg off-axis from the exact backscattering direction). The camera field of view was set to a 1.5-cm-wide square, with the collected light spot in its center. Fig. 1Schematic of the polarimetric imaging setup. P1 and P2 are polarizers; WP1 and WP2 are removable quarter waveplates; L1 and L2 are lenses. In transmission mode, the detection arm is collinear with the incident beam. In the backscattering geometry, the analyzer arm is positioned off the exact retro-reflection direction. 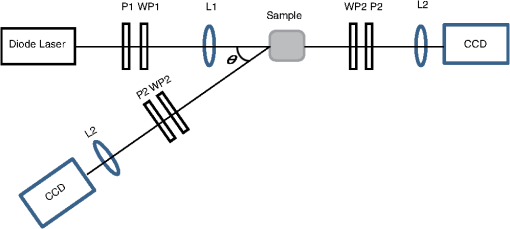 To characterize the interaction of polarized light with a sample of tissue for a specific measurement geometry, Mueller algebra was employed. In Mueller formalism, a polarized light beam is described by the Stokes vector , and its interactions are described by multiplication by a 16-element square () matrix. This matrix, called the Mueller matrix, represents a particular interacting entity (polarizer, wave plate, tissue sample, etc.). If the incident light beam is represented by the Stokes vector , and the output light beam vector is , then the Mueller matrix acts as the polarization transfer function, such that . To calculate the Mueller matrix for a particular tissue sample at particular detection geometry, we performed 24 sequential measurements. For each of four input polarizations [ (linear at 0 deg), (linear at 90 deg), (linear at 45 deg), and (right circular)], intensity profiles at six output polarizations (linear at 0 deg), (linear at 90 deg), (linear at 45 deg), (linear at ), (right circular), and (left circular)] were measured. From these 24 measurements, Mueller matrices for each sample for that specific geometry (transmission/backscattering) were constructed, as described previously in several studies (e.g., Wallenburg et al.26) A complicating issue in turbid polarimetry is the simultaneous occurrence of several polarized light-tissue interactions. This results in complicated cross-talk within and between the Mueller matrix elements and obscures the contribution of the individual sample polarization properties. Several mitigating strategies have been developed recently to deal with this problem.27–30 One approach is termed polar decomposition, originally proposed by Lu-Chipman and adapted by our group for tissue polarimetry.31–34 This decomposition method allows for extraction of individual properties of interest (birefringence, optical activity, depolarization, and diattenuation) from a single measured Mueller matrix.31–34 We have validated this approach using polarized-light Monte-Carlo simulations and phantoms with controlled polarization properties, as well as ex vivo and in vivo tissues.8–10,35 In this method, the tissue Mueller matrix is decomposed into three constituent basis matrices—the depolarization matrix , the retardance matrix , and the diattenuation matrix :27,31–35 This multiplication order (or its reverse) will lead to a physically realizable decomposition in which the basis matrices describe the polarization properties of the sample for the specific geometry.36 For instance, from the retardance matrix, one can calculate a retardance value (proportional to the linear birefringence) and the optical rotation (proportional to circular birefringence) that light experiences upon its interaction with tissue. Note that for different measurement geometries, the effective measurement sampling volumes (and hence the derived polarization characteristics) will be different. The focus of this paper is on the depolarization metrics, which can be calculated from the depolarization matrix . Ignoring for the moment the other two derived basis matrices and the information that they contain, the depolarization matrix can be written in the following form: where is the polarizance vector and is a symmetric submatrix. The diagonal elements of describe the preserved portion of each of the polarization states in the Stokes vector of the light beam. The net depolarization factor, also called the depolarization power, can be defined as 27,33 where , , and are the diagonal elements of . , here termed the total depolarization, yields a quantitative average measure of how a material with this particular Mueller matrix depolarizes light. One can go further and define the linear depolarization, , and circular depolarization, , as follows: Linear depolarization is a measure of the average depolarization of the and components (linear polarization at 0 to 90 deg and 45 to 135 deg) of the Stokes vector interacting with the sample. Correspondingly, circular depolarization is a measure of the depolarization of the component (circular polarization) of the Stokes vector. Using the resultant linear, circular, and total depolarizations allows quantitative comparison of the depolarizing behaviors of different biological tissues at a specific geometry, independent of the illumination polarization state (incident Stokes vector).It is important to distinguish between depolarization metrics derived from the Mueller matrix of the sample and the degree of polarization calculated from the Stokes vector of the light. The former are intrinsic properties of the transfer function of the sample in a particular geometry (as embodied in the tissue Mueller matrix), whereas the latter depend on the sample and on the Stokes vector of the incident light. The derived Mueller metrics thus are more specific to the tissue being studied in that they factor out the confounding effects of the measurement system (incident light polarization). Further, the polar decomposition methods discussed previously permit one to isolate the depolarization effects from other polarizing interactions, something that is not possible with the Stokes descriptors alone. Mathematically, the total degree of polarization in the Stokes description is always higher than the linear and circular degrees of polarizations; an analogous relationship does not hold true for the Mueller matrixderived depolarizations [Eq. (3) through Eq. (5)]. For the remainder of this paper, we discuss depolarization (, , ) as derived from the decomposed Mueller matrices. Following the decomposition procedure, we quantify the intrinsic depolarization parameters of the tissues in each geometry, as derived from Mueller algebra (independent of the incident light polarization state and other tissue polarizing interactions, as per the previous discussion). To gauge the uncertainty in the entire polarimetry methodology, a given Mueller matrix (each with 24 measurements) was recorded three times for all samples in both geometries. Total, linear, and circular depolarization metrics reported in this paper are the average results of these measurements. 3.Results and DiscussionThe measured optical properties (, ) of the swine tissues at 635 nm are shown in Table 1. The reduced scattering coefficient contributes to the extent of photon diffusion in tissue. It is related to the scattering coefficient and anisotropy factor —the mean cosine of the scattering angle-through the relation . The transport albedo is a dimensionless quantity defined as . In addition to their role in interpreting tissue polarimetry results, these measured optical properties may contribute to recent tissue optics compilations.37–40 Polarized light imaging allows the construction of spatial maps in which the Mueller matrix derived parameters may be plotted. In Fig. 2, we illustrate the calculated total, linear, and circular depolarization of kidney cortex samples in the transmission and backscattering geometries. The data shown in each map is the average of three independent measurements. The average standard deviation over all the pixels in the image is . This number is representative of the average uncertainty in the derived polarization metrics, characteristic of the system noise over the measurement period. Fig. 2Depolarization maps of the kidney cortex derived from the polar decomposition of the measured Mueller matrices. The columns indicate the total, linear, and circular depolarizations, and the rows indicate the different detection geometries and sample thickness. The and axes indicate the length and width, respectively. The color bar indicates depolarization levels as specified on the percent scale, where dark blue signifies regions where light remains most polarized and deep red indicates regions where light has lost most of its polarization. The error bar size is equal to the line thickness. 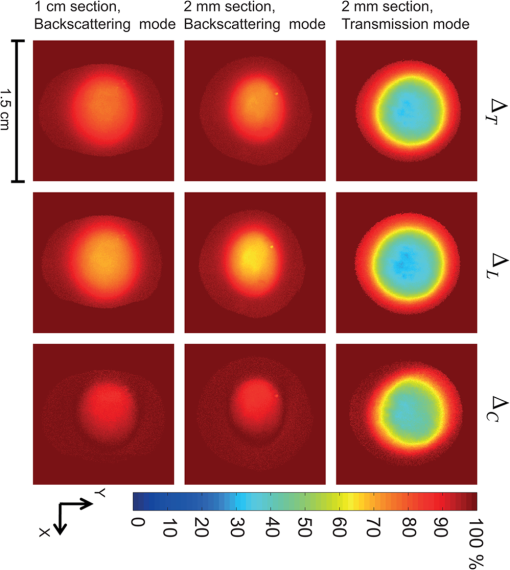 Table 1Measured optical properties of several swine tissues at 635 nm.
Several prominent features are visually evident from Fig. 2. First, linear depolarization is lower than circular depolarization for both transmission and backscattering geometries, as well as for both 2-mm and 1-cm kidney cortex samples. In other words, linear polarization is better preserved than circular polarization in this tissue type. Second, the total depolarization is observed to be lowest in transmission mode for 2-mm samples and greatest in the backscattering mode for the 1-cm sample. Such images have been obtained for all the examined tissues; to enable quantitative analysis and inter-tissue comparisons, we averaged the data along a 0.1-cm central strip in the direction, and plotted the total depolarization along the direction, as shown in Fig. 3. Fig. 3Depolarization of different tissues’ samples in the axial direction. (a) Total depolarization in transmission mode for the 2-mm samples; (b) total depolarization in backscattering mode for 2-mm samples; (c) total depolarization in backscattering mode for 1-cm samples. Brain and tendon are not shown in (a) because they exhibit total depolarization greater than 98.5% in transmission mode, which is in the range of the system noise level. 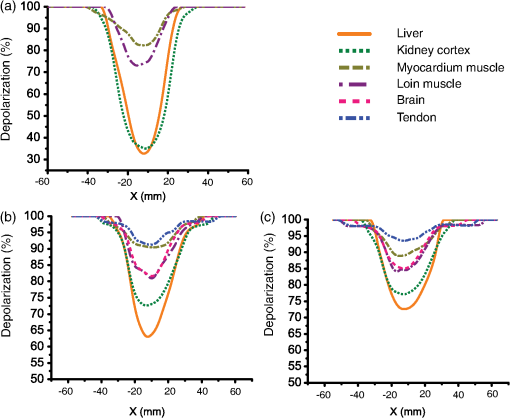 Comparing panels (a) and (b) of Fig. 3, one can conclude that all the tissues except tendon and brain are less depolarizing in transmission geometry rather than in backscattering. For brain and tendon, the transmission mode is not as useful as depolarization becomes too severe (greater than 98.5% even for this relatively thin, 2-mm slab) and essentially all polarized light information is lost. Measurements with such large depolarization are not reliable and lie in the range of the polarimetry system’s noise level. The transmission geometry thus becomes untenable for tissue polarimetry as either physical thickness or optical properties increase sufficiently. Hence, the brain and tendon results are not included in Fig. 3(a). Such extensive depolarization is likely because brain and tendon both have transport albedos higher than 0.97 (Table 1). A large transport albedo signifies the predominance of multiple scattering, with its variety of long, zigzag paths that effectively depolarize the light.41,42 Furthermore, scattering in biological tissues is generally forward-peaked, with the anisotropy factor varying between 0.9 and 0.99.19 A large transport albedo and forward-peaked scattering implies that photons detected in backscattering geometry have experienced fewer scattering events and on average have traveled shorter path-lengths compared to the photons detected in transmission geometry. In fact, most photons detected in the reflection mode in these types of scattering media have sampled mostly near-surface layers, as we have recently demonstrated by Monte Carlo modeling studies.42–44 As a result, less depolarization is observed for high transport albedo tissues in backscattering mode. A general trend in the total depolarization values of the tissues emerges from a closer examination of Fig. 3. Liver has the minimum total depolarization in all measurement geometries, whereas tendon appears to have the highest total depolarization among all tissue types. The tissues arranged in the order of increasing depolarization are liver, kidney, loin muscle, brain, myocardium muscle, and tendon. Comparing Fig. 3 and Table 1, one notices a correlation between total depolarization rates and the transport albedos of the corresponding tissue type. To explore this further, the minimums of total, linear, and circular depolarization (Fig. 3) are plotted against the tissues’ transport albedos (Table 1) in Fig. 4. As previously mentioned, brain and tendon data in transmission mode are not shown because they exhibit depolarizations . Analysis of Fig. 4 reveals that tissues with higher transport albedos are generally more depolarizing. This trend appears to hold for all tissues, detection geometries, and sample thicknesses. The relations’ depolarization and transport albedo have been quantified with the correlation coefficient (r) between them, as shown in Fig. 4. These correlations include both anisotropic and isotropic tissues; as seen, anisotropic tissues shows higher depolarization compared to the isotropic one with the same transport albedo. For example, myocardium muscle, whose is not exceptionally high (close to the value for kidney cortex), exhibits very high total depolarization. In addition, when comparing tendon tissue depolarization to that of the brain, tendon exhibits a higher depolarization despite having a comparable . One possible reason for both of these observations could arise from the intrinsic anisotropy exhibited by the different tissue types. Myocardium muscle and tendon (and loin muscle, to a lesser extent) are known to be highly anisotropic relative to the other tissues.17 The underlying birefringence that causes this tissue anisotropy thus may contribute to depolarization over and above the transport albedo effect invoked previously. Specifically in tissues, birefringence magnitude and orientation may be spatially inhomogeneous, changing in different regions of tissues/microdomains;34,35,45,46 in these regions, polarized light undergoes additional randomization, and therefore the total depolarization increases.15,45 Note that we specifically concentrate on depolarization phenomena here and do not analyze the derived birefringence magnitude and orientation (derived from the retardance matrix ), which are important in anisotropic tissues such as skeletal muscle;33,46 an ongoing study examining birefringence phenomena in greater detail (e.g., the effects of variable spatial domains of different magnitude/orientation of birefringence) will be reported elsewhere. Fig. 4Depolarization of different tissues related to their transport albedos. Liver is indicated by L; kidney, myocardium muscle, loin muscle, brain, and tendon are indicated by K, M, H, B, and T, respectively. (a), (b), and (c) show total, linear, and circular depolarizations in transmission mode for the 2-mm samples. (d), (e), and (f) show total, linear, and circular depolarizations in backscattering mode for the 2-mm samples. (g), (h), and (i) show total, linear, and circular depolarization for the 1-cm samples in the backscattering mode. , the correlation coefficient between the depolarization and the transport albedo for each graph, is also shown. The correlation coefficient (Pearson coefficient) indicates how well data fits a straight line, and it is calculated as for data points, with and and being their mean values. In this case, and . Note that brain and tendon data is not included in (a)–(c) owing to their high depolarization (). 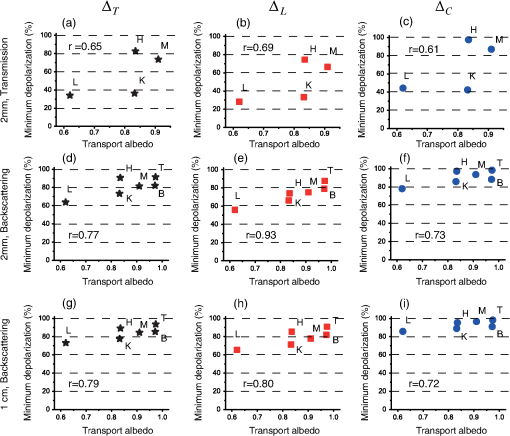 Another interesting trend evident from Fig. 4 is the preferential retention of linear polarization states compared to circular polarization states, in all tissues, for both geometries. Interestingly, the correlation coefficient values are also higher in cases of linear depolarization. Better preservation of linear polarization is usually associated with Rayleigh-like scattering, whereas tissues are often considered Mie-like in their scattering behavior. Thus, identifying the appropriate dominant scattering regime in biological tissues is tricky. Conventional wisdom is that tissues are composed of large scatterers and thus exhibit Mie-like scattering behavior, consistent with the predominantly forward-peaked nature of the observed scattering phase function (high values of , as discussed previously). While cells and subcellular organelles such as the nucleus and mitochondria are within the Mie scattering regime, extracellular matrix elements such as collagen fibrils fall in the Rayleigh scattering regime.17 In suspensions containing both small and large scatterers, Ghosh et al. have shown that scattering behavior is dominated by smaller particles.47 This was also observed in simulations whereby small organelles were shown to contribute heavily to light scattering in cells.48 On the basis of size alone, one can therefore expect that biological tissues may be described better by the Rayleigh regime, with an alternate explanation for the high observed -values. The other important determining factor is the relative refractive index. Based on another study by Ghosh et al., if a phantom has large scatterers and a high anisotropy factor but a small relative refractive index contrast, it does not fall in the Mie scattering regime (e.g., circular polarization is no longer preserved better than linear, as it should in the “classical” Mie regime).49 Relative refractive index contrast and scatterer size thus are both important in determining the dominant scattering behavior. Interestingly, Sankaran et al. also reported better retention of linear over circular polarization states in transmission, measured though ∼-thick slabs of tissues 15 and through dense suspensions of high-contrast tissue simulating phantoms (compared to dilute phantoms);50 for the latter, they invoked the mechanisms of correlated (dependent) scattering engendered by dense packing to explain their findings. The preferential retention of linear polarization states observed in our study is also indicative of Rayleigh scattering, although the exact mechanism remains unclear. Given the complicated nature of tissue scattering, likely both small scatterers and large scatterers (with small relative refractive index and large anisotropy factor g), augmented by compactness, are contributing. It is also worth mentioning that in all these studies, linear and circular degrees of polarization were based on the Stokes parameters of the light beam and thus depend somewhat on the input light parameters and are unable to account for other tissue polarizing effects; in contrast, our reported results are derived from the tissue Mueller matrix via polar decomposition and thus are free of these two confounding effects. Despite these differences, both approaches suggest the preferred preservation of linear polarization states over circular polarization states in biological tissues. We are currently conducting a controlled phantom study to explore further the linear and circular depolarization behavior as a method to ascertain the dominant scattering regime and the essential medium characteristics determining it.51 4.ConclusionsIn summary, we have measured depolarization and optical properties in a variety of biological tissues, and we have quantified the correlations between them. We used polarized light imaging and Mueller matrix polar decomposition to extract linear, circular, and total depolarizations in several freshly excised bulk swine tissues (liver, tendon, kidney cortex, brain, myocardium muscle, and loin muscle). Both transmission and reflection geometries were used to examine 2-mm and 1-cm tissue slabs. Bulk optical properties of the different tissues were measured and related to the derived depolarization behavior. A linear correlation between transport albedo and depolarization was observed and quantified. Furthermore, in highly anisotropic tissues like cardiac muscle and tendon, birefringence increases depolarization over and above the multiple scattering effects, as determined by the high transport albedo (i.e., in spite of the relatively low of these birefringent tissues). Also observed was the preferential preservation of linear over circular polarized light in all tissue types and for all detection geometries. A tentative explanation invoking effective scatterer size and relative refractive index contrast was provided. Finally, for high-transport albedo tissues such as the brain and tendon, light experiences more depolarization through propagation in the forward direction (transmission mode) compared to the backscattering direction. In fact, transmission detection geometry becomes untenable for polarimetry applications for even 2-mm samples of brain and tendon because of extensive polarization loss. Detailed studies of depolarization behavior in various bulk animal tissues should help researchers understand the underlying biophysics of polarized light-tissue interactions, provide polarization properties of various tissue types that are generally unavailable in the literature, and assist in experimental design for polarized light imaging and for polarimetry-based tissue characterization studies. AcknowledgmentsThis work was supported by the Natural Sciences and Engineering Research Council of Canada. Manzoor Ahmad acknowledges the Higher Education Commission (HEC), Islamabad, Pakistan, for the financial support through the Indigenous Fellowship Scheme for PhD studies. ReferencesR. K. CurtisD. R. Tyson,
“Birefringence: polarization microscopy as a quantitative technique of human hair analysis,”
J. Soc. Cosmet. Chem., 27 411
–431
(1976). JSCCA5 Google Scholar
R. Oldenbourg,
“A new view on polarization microscopy,”
Nature, 381 811
–812
(1996). http://dx.doi.org/10.1038/381811a0 NATUAS 0028-0836 Google Scholar
W. Groneret al.,
“Orthogonal polarization spectral imaging: a new method for study of the microcirculation,”
Nat. Med., 5
(10), 1209
–1212
(1999). 1078-8956 Google Scholar
X. GanS. P. SchildersM. Gu,
“Image enhancement through turbid media under a microscope by use of polarization gating methods,”
J Opt. Soc. Am. A, 16
(9), 2177
–2184
(1999). http://dx.doi.org/10.1364/JOSAA.16.002177 1084-7529 Google Scholar
M. R. Scheinfeinet al.,
“Scanning electron microscopy with polarization analysis (SEMPA),”
Rev. Sci. Instrum., 61
(10), 2501
–2527
(1990). http://dx.doi.org/10.1063/1.1141908 RSINAK 0034-6748 Google Scholar
W. E. MoernerD. P. Fromm,
“Methods of single-molecule fluorescence spectroscopy and microscopy,”
Rev. Sci. Instrum., 74
(8), 3597
–3619
(2003). http://dx.doi.org/10.1063/1.1589587 RSINAK 0034-6748 Google Scholar
E. J. G. PetermanH. SosaW. E. Moerner,
“Single-molecule fluorescence spectroscopy and microscopy of biomolecular,”
Annu. Rev. Phys. Chem., 55 79
–96
(2004). http://dx.doi.org/10.1146/annurev.physchem.55.091602.094340 ARPLAP 0066-426X Google Scholar
M. F. G. Woodet al.,
“Polarization birefringence measurements for characterizing the myocardium, including healthy, infarcted, and stem-cell-regenerated tissues,”
J. Biomed. Opt., 15
(4), 047009
(2010). http://dx.doi.org/10.1117/1.3469844 JBOPFO 1083-3668 Google Scholar
M. F. G. Woodet al.,
“Proof-of-principle demonstration of a Mueller matrix decomposition method for polarized light tissue characterization in vivo,”
J. Biomed. Opt., 14 014029
(2009). http://dx.doi.org/10.1117/1.3065545 JBOPFO 1083-3668 Google Scholar
M. F. G. WoodX. GuoI. A. Vitkin,
“Polarized light propagation in multiply scattering media exhibiting both linear birefringence and optical activity: Monte Carlo model and experimental methodology,”
J. Biomed. Opt., 12 014029
(2007). http://dx.doi.org/10.1117/1.2434980 JBOPFO 1083-3668 Google Scholar
S. G. Demoset al.,
“Time-resolved degree of polarization for human breast tissue,”
Opt. Commun., 124 439
–442
(1996). http://dx.doi.org/10.1016/0030-4018(95)00693-1 OPCOB8 0030-4018 Google Scholar
S. L. JacquesJ. R. RomanK. Lee,
“Imaging superficial tissues with polarized light,”
Laser. Surg. Med., 26 119
–129
(2000). http://dx.doi.org/10.1002/(ISSN)1096-9101 LSMEDI 0196-8092 Google Scholar
S. L. JacquesJ. C. Ramella-RomanK. Lee,
“Imaging skin pathology with polarized light,”
J. Biomed. Opt., 7
(3), 329
–340
(2002). http://dx.doi.org/10.1117/1.1484498 JBOPFO 1083-3668 Google Scholar
I. A. VitkinR. C. N. Studinski,
“Polarization preservation in diffusive scattering from in vivo turbid biological media: effects of tissue optical absorption in the exact backscattering direction,”
Opt. Commun., 190 37
–43
(2001). http://dx.doi.org/10.1016/S0030-4018(01)01080-X OPCOB8 0030-4018 Google Scholar
V. Sankaran Jr.J. T. WalshD. J. Maitland,
“Comparative study of polarized light propagation in biologic tissues,”
J. Biomed. Opt., 7
(3), 300
–306
(2002). http://dx.doi.org/10.1117/1.1483318 JBOPFO 1083-3668 Google Scholar
M. Antonelliet al.,
“Mueller matrix imaging of human colon tissue for cancer diagnostics: how Monte Carlo modeling can help in the interpretation of experimental data,”
Opt. Express, 18
(10), 10200
–10208
(2010). http://dx.doi.org/10.1364/OE.18.010200 OPEXFF 1094-4087 Google Scholar
V. Tuchin, Tissue Optics., 2d ed.SPIE, Washington D. C.
(2007). Google Scholar
M. F. G. Woodet al.,
“Effects of formalin fixation on tissue optical polarization properties,”
Phys. Med. Biol., 56 N115
–N122
(2011). http://dx.doi.org/10.1088/0031-9155/56/8/N01 PHMBA7 0031-9155 Google Scholar
G. JarryF. HenryR. Kaiser,
“Anisotropy and multiple scattering in thick mammalian tissues,”
J. Opt. Soc. Am. A, 17
(1), 149
–53
(2000). http://dx.doi.org/10.1364/JOSAA.17.000149 1084-7529 Google Scholar
M. Notohamiprodjoet al.,
“Diffusion tensor imaging of the kidney with parallel imaging: initial clinical experience,”
Investig. Radiol., 43
(10), 677
–685
(2008). Google Scholar
R. A. CohenL. H. Sweet, Brain Imaging in Behavioural Medicine and Clinical Neuroscience, Springer, New York
(2010). Google Scholar
D. Jones, Diffusion MRI, Theory, Methods, and Applications, Springer, New York
(2004). Google Scholar
T. J. FarrellM. C. PattersonB. C. Wilson,
“A diffusion theory model of spatially resolved, steady-state diffuse reflectance for the non-invasive determination of tissue optical properties in vivo,”
Med. Phys., 19 879
–888
(1992). http://dx.doi.org/10.1118/1.596777 MPHYA6 0094-2405 Google Scholar
A. KimB. C. Wilson,
“Measurement of ex vivo and in vivo tissue optical properties: methods and theories.,”
In Optical-Thermal Response of Laser-Irradiated Tissues, Springer, New York
(2011). Google Scholar
A. Kimet al.,
“A fiberoptic reflectance probe with multiple source-collector separations to increase the dynamic range of derived tissue optical absorption and scattering coefficients,”
Opt. Express, 18
(6), 5580
–5594
(2010). http://dx.doi.org/10.1364/OE.18.005580 OPEXFF 1094-4087 Google Scholar
M. A. Wallenburget al.,
“Comparison of optical polarimetry and diffusion tensor MR imaging for assessing myocardial anisotropy,”
J. Innov. Opt. Health Sci., 3
(2), 109
–121
(2010). Google Scholar
S. LuR. A. Chipman,
“Interpretation of Mueller matrices based on polar decomposition,”
J. Opt. Soc. Am. A, 13
(5), 1106
–1113
(1996). http://dx.doi.org/10.1364/JOSAA.13.001106 1084-7529 Google Scholar
R. Ossikovski,
“Analysis of depolarizing Mueller matrices through a symmetric decomposition,”
J. Opt. Soc. Am. A, 26
(5), 1109
–1118
(2009). http://dx.doi.org/10.1364/JOSAA.26.001109 1084-7529 Google Scholar
R. Ossikovski,
“Differential matrix formalism for depolarizing anisotropic media,”
Opt. Lett., 36
(12), 2330
–2332
(2011). http://dx.doi.org/10.1364/OL.36.002330 OPLEDP 0146-9592 Google Scholar
N. Ortega-QuijanoJ. L. Arce-Diego,
“Mueller matrix differential decomposition,”
Opt. Lett., 36
(10), 1942
–1944
(2011). http://dx.doi.org/10.1364/OL.36.001942 OPLEDP 0146-9592 Google Scholar
M. K. Swamiet al.,
“Polar decomposition of 3×3 Mueller matrix: a tool for quantitative tissue polarimetry,”
Opt. Express, 14
(20), 9324
–9337
(2006). http://dx.doi.org/10.1364/OE.14.009324 OPEXFF 1094-4087 Google Scholar
M. F. G. WoodN. GhoshI. A. Vitkin,
“Mueller matrix decomposition for extraction of individual polarization parameters from complex turbid media exhibiting multiple scattering, optical activity, and linear birefringence,”
J. Biomed. Opt., 13
(4), 044036
(2008). http://dx.doi.org/10.1117/1.2968198 JBOPFO 1083-3668 Google Scholar
N. Ghoshet al.,
“Mueller matrix decomposition for polarized light assessment of biological tissues,”
J. Biophoton., 2
(3), 145
–156
(2009). http://dx.doi.org/10.1002/jbio.v2:3 JBOIBX 1864-063X Google Scholar
X. LiG. Yao,
“Mueller matrix decomposition of diffuse reflectance imaging in skeletal muscle,”
Appl. Opt., 48
(14), 2625
–2631
(2009). http://dx.doi.org/10.1364/AO.48.002625 APOPAI 0003-6935 Google Scholar
N. GhoshM. F. G. WoodI. A. Alex Vitkin,
“Polarimetry in turbid, birefringent, optically active media: a Monte Carlo study of Mueller matrix decomposition in the backscattering geometry,”
J. Appl. Phys., 105 102023
(2009). http://dx.doi.org/10.1063/1.3116129 JAPIAU 0021-8979 Google Scholar
N. GhoshM. F. G. WoodI. A. Vitkin,
“Influence of the order of the constituent basis matrices on the Mueller matrix decomposition-derived polarization parameters in complex turbid media such as biological tissues,”
Opt. Commun., 283 1200
–1208
(2010). http://dx.doi.org/10.1016/j.optcom.2009.10.111 OPCOB8 0030-4018 Google Scholar
W. F. CheongS. A. PrahlA. J. Welch,
“A review of the optical properties of biological tissues,”
IEEE J. Quant. Electron., 26
(12), 2166
–2185
(1990). http://dx.doi.org/10.1109/3.64354 IEJQA7 0018-9197 Google Scholar
C. J. HourdakisA. Penis,
“A Monte Carlo estimation of tissue optical properties for use in laser dosimetry,”
Phys. Med. Biol., 40 351
–364
(1995). http://dx.doi.org/10.1088/0031-9155/40/3/002 PHMBA7 0031-9155 Google Scholar
G. JagajothiS. Raghavan,
“Estimation and measurement of biological tissue using optical simulation method,”
Prog. Electromagn. Res. M, 6 155
–165
(2009). http://dx.doi.org/10.2528/PIERM09021604 1070-4698 Google Scholar
R. SrinivasanD. KumarM. Singh,
“Optical characterization and imaging of biological tissues,”
Curr. Sci., 87 218
–227
(2004). CUSCAM 0011-3891 Google Scholar
M. S. PattersonB. C. WilsonD. Wyman,
“The propagation of optical radiation in tissue, II: optical properties of tissues and resulting fluence distributions,”
Laser. Med. Sci., 6
(2), 379
–390
(1991). http://dx.doi.org/10.1007/BF02042460 0268-8921 Google Scholar
X. GuoM. F. G. WoodI. A. Vitkin,
“Monte Carlo study of pathlength distribution of polarized light in turbid media,”
Opt. Express, 15
(3), 1348
–1360
(2007). http://dx.doi.org/10.1364/OE.15.001348 OPEXFF 1094-4087 Google Scholar
X. GuoM. F. G. WoodI. A. Vitkin,
“Angular measurements of light scattered by turbid chiral,”
J. Biomed. Opt., 11
(4), 041105
(2006). http://dx.doi.org/10.1117/1.2339134 JBOPFO 1083-3668 Google Scholar
X. GuoM. F. G. WoodI. A. Vitkin,
“Depolarization of light in turbid media: a scattering event resolved Monte Carlo study,”
Appl. Opt., 49
(2), 153
–162 http://dx.doi.org/10.1364/AO.49.000153 APOPAI 0003-6935 Google Scholar
Y. P. Sinichkinet al.,
“The effect of the optical anisotropy of scattering media on the polarization state of scattered light,”
Optic Spectros., 101
(5), 802
–810
(2006). http://dx.doi.org/10.1134/S0030400X06110221 OPSUA3 0030-400X Google Scholar
D. A. ZimnyakovY. P. SinichkinO. V. Ushakova,
“Optical anisotropy of fibrous biological tissues: analysis of the influence of structural properties,”
Quant. Electron., 37
(8), 777
–783
(2007). http://dx.doi.org/10.1070/QE2007v037n08ABEH013611 QUELEZ 1063-7818 Google Scholar
N. Ghoshet al.,
“Anomalous behavior of depolarization of light in a turbid medium,”
Phys. Lett. A., 354 236
–242
(2006). http://dx.doi.org/10.1016/j.physleta.2006.01.053 PYLAAG 0375-9601 Google Scholar
A. DunnR. Richards-Kortum,
“Three-dimensional computation of light scattering from cells,”
IEEE J. Sel. Top. Quant. Electron., 2
(4), 898
–905
(1996). http://dx.doi.org/10.1109/2944.577313 IJSQEN 1077-260X] Google Scholar
N. Ghoshet al.,
“Depolarization of light in a multiply scattering medium: effect of the refractive index of a scatterer,”
Phys. Rev. E, 70 066607
(2004). http://dx.doi.org/10.1103/PhysRevE.70.066607 PLEEE8 1539-3755 Google Scholar
V. SankaranJ. T. WalshD. J. Maitland,
“Polarized light propagation through tissue phantoms containing densely packed scatterers,”
Opt. Lett., 25
(4), 239
–241
(2000). http://dx.doi.org/10.1364/OL.25.000239 OPLEDP 0146-9592 Google Scholar
M. Ahmadet al.,
“Do different turbid media with matched bulk optical properties also exhibit similar polarization properties?,”
Biomed. Opt. Express, 2
(12), 3248
–3258
(2011). http://dx.doi.org/10.1364/BOE.2.003248 BOEICL 2156-7085 Google Scholar
|

