|
|
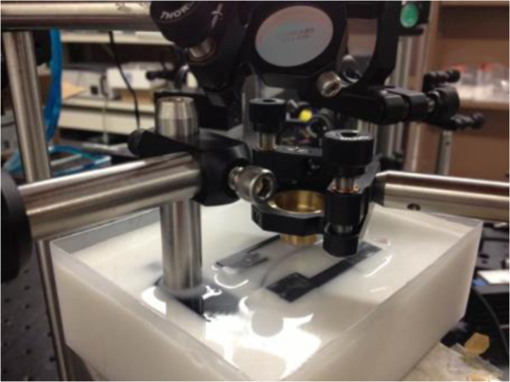
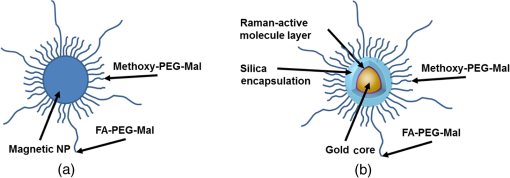
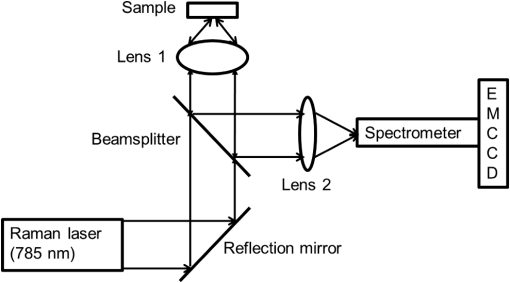
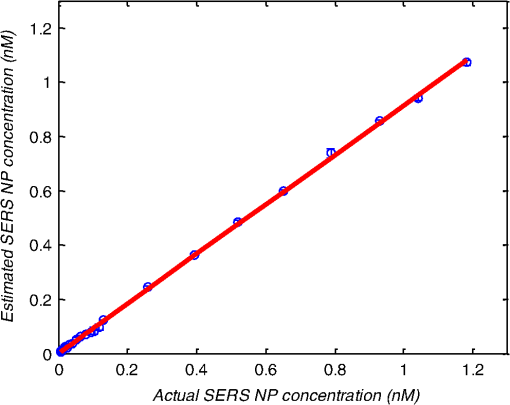
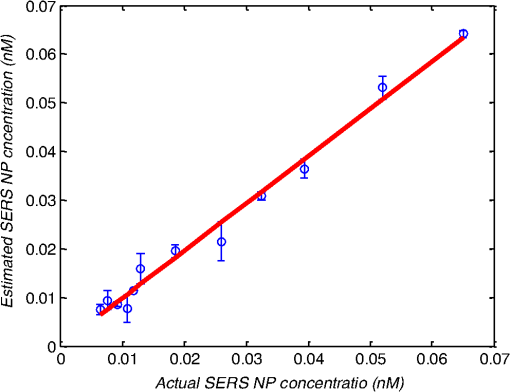
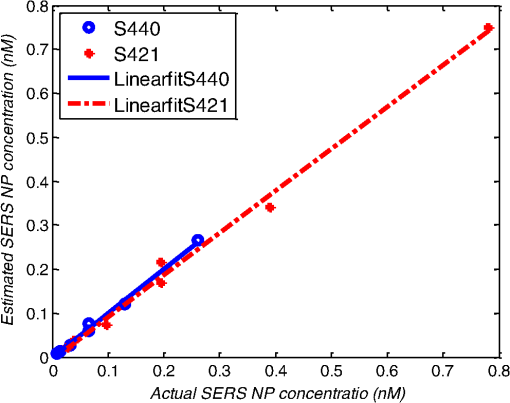
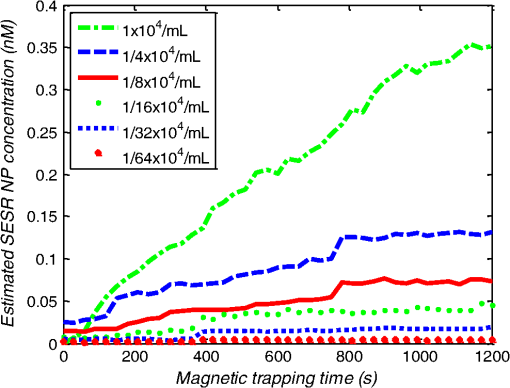
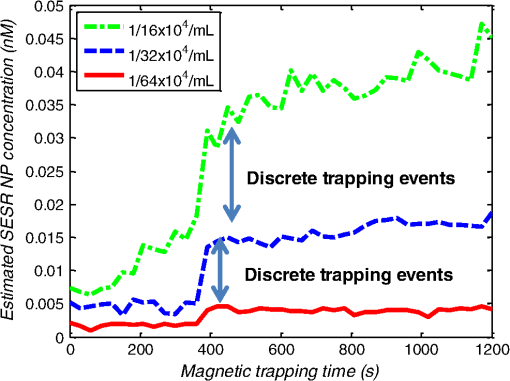
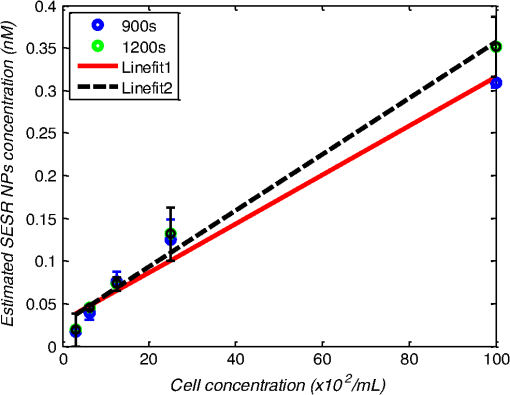
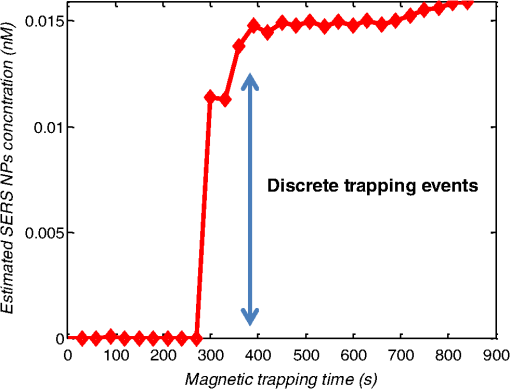
1.IntroductionMetastasis, the spread of cancer cells from the primary tumor to distant sites in the body, is known to cause of cancer-associated deaths.1 Circulating tumor cells (CTCs) are a marker of cancer metastasis and hence a source of biomarkers for tumor research and diagnosis. Detecting CTCs helps in diagnosis at the early stages of metastasis and may have important prognostic and therapeutic implications.2 However, directly identifying CTCs is difficult due to their low concentration in blood, which may be on the order of 1 to compared to millions of white blood cells and nearly a billion red blood cells per mL of blood.3 Both capture sensitivity and specificity are critical for the detection of CTCs. A variety of enrichment technologies have been developed to separate and identify CTCs.4–7 Messenger RNA-based strategies with PCR amplification do not allow collection of cells for further analysis and may miss CTCs that exhibit unexpected phenotypes. In immunomagnetic methods, iron or magnetic beads coated with monoclonal antibodies specific to cell surface oncoantigens or cell-specific cytokeratin and common leukocyte antigen (CD45) are used to capture and enrich CTCs.8 Methods including flow cytometry are used for enumeration of CTCs in whole blood and have high sensitivity and reproducibility, but rely on expensive specific equipment. Additionally, high specificity often requires multiple fluorescent labels to avoid false positives, yet spectral overlap of absorption and emission spectra from fluorescent labels can limit multiplexing and hence specificity. Additionally, background autofluorescence can significantly limit detection sensitivity in tissue. Microfluidic separation alone or in combination with immunomagnetic assays has also been used to capture and analyze CTCs.9,10 Filter-based physical methods have been developed to enable capture of CTCs based on size, keeping the CTCs while filtering out blood cells.11 However, the above methods are not suitable for in vivo detection or enrichment. In 2009, Galanzha et al.12 reported the rapid photoacoustic detection of magnetically captured CTCs in vivo using magnetic and optically absorbing nanoparticles. In 2012, Wei et al.13 demonstrated the magnetomotive photoacoustic imaging of polystyrene beads using magnetic and photoacoustic contrast agents. Photoacoustic flow cytometry approaches have also been used to sort and detect CTCs.14 However, these photoacoustic approaches have, thus far, demonstrated the limited multiplexing capacity to improve the specificity of detection. Raman spectroscopy presents high multiplexing capabilities due to the multiple narrow linewidths of spectral peaks. Surface-enhanced Raman scattering nanoparticles (SERS NPs) produce strong Raman scattering optical signatures unique to each “flavor” of SERS NPs, and offer a way of spectral fingerprinting that may provide extremely high specificity when targeting multiple cell biomarkers. In 2009, Zavaleta et al.15 demonstrated the ability to produce multiplexed in vivo images with 10 species of SERS NPs at picomolar (pM) sensitivities with ultralow background levels. In 2011, Wang et al.16 reported using SERS NPs to detect CTCs in human peripheral blood. However, their method lacked an enrichment technique. In 2010, Jun et al. demonstrated that silica-encapsulated magnetic NPs (M-SERS dots) produce strong SERS signals and have magnetic properties that can be used for cancer-cell targeting and separation.17 In 2014, Quaresma et al. reported that star-shaped gold-coated magnetic nanoparticles (MNPs) with unique properties demonstrated SERS detection of Astra Blue, and effective magnetic separation of a histidine-tagged maltose binding protein from a crude cell-extract.18 However, neither of these studies examined the magnetic trapping of CTCs. Additionally, previously developed magnetic-SERS NPs could produce false-positive signals during magnetic trapping when the intention is to detect only targeted CTCs. To achieve detection of CTCs with high sensitivity and specificity, we propose a combination of magnetic trapping and multiplex detection of CTCs by using targeted magnetic NPs and SERS probes. We demonstrate an SERS imaging system with 1-pM sensitivity, 2.5-mm penetration depth, and high linearity between estimated SERS NP concentrations and actual SERS NP concentrations. Our magnetic trapping system is able to effectively trap cells at flow velocities ranging from 0.2 to . Currently, both SERS NPs and MNPs are targeted to folate receptor, overexpressed on many cancer cells but absent on most normal blood cells. We show that only cells targeted with both SERS NPs and MNPs exhibit an increasing SERS signal due to magnetic accumulation of CTCs. Hence, the SERS signal increase correlates with CTC detection. Free NPs or CTCs targeted with only SERS NPs or only MNPs do not exhibit this optical increase. In addition, discrete SERS signals were detected in the magnetic trapping zone from a mixture of flowing dual-nanoparticle labeled HeLa cells and phosphate buffered saline (PBS) or rat blood indicative of single cell trapping events. This technology could be used for in vitro or in vivo detection of CTCs, and may prove important for cancer research, molecular diagnostics, and personal medicine. 2.Materials and Methods2.1.Folate Receptor TargetingCompared to low levels of folate receptors in most normal cells, many cancer cells (breast, lung, kidney, ovary, colon, brain, and myelogenous leukemia) significantly overexpress folate receptors to facilitate rapid cell division.19 We conjugate nanoparticles with folate to target folate receptors of cancer cells. 2.2.Magnetic NanoparticlesMNPs (screenMAG-Thiol, chemicell™ GmbH, Berlin, Germany) consisted of 500-nm-diameter magnetic fluorescent silica particles with nearly 1 million SH-groups per particle. To target folate receptors, thiolated MNPs (1.25 nM, ) were added to a solution of folate-PEG5k-maleimide (Nanocs Inc., New York, PG2-FAML-5k, 0.8 mM) and methoxy-PEG2k-maleimide (Sigma™ Aldrich, Oakville, Canada, 4 mM) and mixed at room temperature by a rocker overnight. Excess folate-PEG-maleimide was separated from MNPs by three runs of centrifugation (14,000 rpm, 5 min), and resuspension in 2-(N-morpholino)ethanesulfonic acid (MES, Fisher Science) buffer. The diagrammatic representation of the conjugated MNP is shown in Fig. 1(a). 2.3.Surface-Enhanced Raman Scattering NanoparticlesSERS NPs (S440 and S420, Oxonica Materials Inc., Mountain View, California) are composed of a 60-nm-diameter Au core with a Raman active molecular monolayer adsorbed onto it. They are encapsulated with a 30-nm silica shell, with an overall diameter of 120 nm. Each type of SERS NP consists of a different Raman active molecular layer with its own unique Raman spectrum. To simplify, instead of nomenclature based on the Raman active molecular monolayer, a three-digit suffix (e.g., S440, S421) is used to name each type of SERS NP. Using a similar procedure as MNPs targeting, a solution of folate-PEG5k-maleimide (Nanocs Inc., PG2-FAML-5k, 0.8 mM) and methoxy-PEG2k-maleimide (Sigma Aldrich, 4 mM) was added to thiolated SERS NPs (1.3 nM, ) and mixed at room temperature on a rocker overnight, followed by three runs of centrifugation (14,000 rpm, 5 min) to eliminate excess folate-PEG-maleimide and resuspension in MES buffer. The diagrammatic representation of the conjugated SERS NP is shown in Fig. 1(b). 2.4.Magnetic Trapping SystemTo achieve efficient magnetic trapping, two rectangular neodymium rare earth magnets (, Indigo™ Instruments, Waterloo, Canada) separated by a distance of 20 mm were used to provide a 0.07 T external magnetizing field in the tubing area. This was used to polarize the MNPs for subsequent efficient trapping. A 1-in. cone magnet (Cone-Cone0100N, SuperMagnetMan) pointed at the optical focus was used for generating a high magnetic field gradient for trapping. Cells were flowed inside Tygon™ tubing of 1-mm inner diameter controlled by a syringe pump (NE-300, New Era Pump Systems, Inc., Farmingdale). 2.5.Raman Imaging SystemOur Raman imaging setup (Fig. 2) uses a 785-nm Raman laser (FB-785-350-FS-FS-1-1, RGBLase LLC, Fremont, California) as an excitation source. This laser provides up to 350 mW (while the power used in our experiment was 54 mW) and has a 20-dB spectral linewidth of 0.4 nm and side lobe suppression ratio . The narrow excitation linewidth is important since Raman peaks have linewidths of only a couple of nanometers or less. Raman spectra were captured using a custom Raman imaging spectrometer (RASPPEC-785-HR, P&P Optica Inc. Kitchener, Ontario, Canada). The imaging spectrometer is capable of measuring the Raman spectra from each point in a line. The input aperture is a slit of size . Light from this slit is collimated, passed through a gel grating, then refocused onto a two-dimensional camera sensor array. The imaging spectrometer presents a 21-deg bend to ensure collection of diffracted light in the desired spectral range. The gel-grating is a through-transmission grating with offering a spectral range of 815 to 895 nm. The imaging spectrometer offers a spectral resolution of with an f# of 3.5. A 785-nm ultra-steep long-bandpass cutoff (OD 8) is used as Raman filter. Spectra are detected using a liquid-cooled electron-multiplying (EM) CCD camera (DU971N-UV, Andor Technology, South Windsor). This camera has dark current noise as low as at and a quantum efficiency of at peak wavelengths, in the measurement wavelength range. The sensor head is a front-illuminated EMCCD array. The camera is mounted such that the spectra are recorded along the short axis of the sensor. The camera can be rotated 90-deg relative to the spectrometer if needed. Pixel elements are . Currently, we use only a few spatial channels ( vertical pixels) of our imaging spectrometer system for point illumination and collection. The EM gain was set at 10, integration time was 3 s, and sensor temperature was liquid-cooled to . An aspheric lens (A240TM-B, Thorlabs, Newton) with a 0.5 numerical aperture (NA) and 8-mm focal length (shown as lens 1 in Fig. 2) was used for both illumination and light collection. Collimated backscattered light from the sample was redirected via a beam splitter (DMSP805L, Thorlabs) and focused into the input aperture of the imaging spectrometer via a 0.3-NA lens (AC127-019-B-ML, Thorlabs) with a magnification of . Algorithms to correct for spherical aberrations on the camera sensor were provided by the spectrometer manufacturer. Demixing was based on a classical linear least-squares method with an optional positivity constraint. In experiments, background noise and fluorescence signals generated from real tissue should be acquired before SERS detection and used as demixing factors to eliminate interference effects. Sensitivity was evaluated by detecting SERS signals from serial dilutions of SERS NP solutions in PCR tubes. 3.Experiments and Results3.1.SensitivityThe data present a highly linear relationship between estimated SERS NP concentrations and actual SERS NP concentrations with (Fig. 3). Figure 4 is a close-up view of the low concentration part in Fig. 3. It can be seen that the high linearity was maintained for SERS NP concentrations even below 10 pM. Defining sensitivity as the concentration of SERS NPs detectable with a unity signal-to-noise ratio, our sensitivity is estimated as . To clarify, when reporting nanoparticle concentrations, we report the number density of nanoparticles and these concentrations should not be confused with the effective Raman reporter concentrations. To estimate the maximum depth of penetration of our system for possible future in vivo applications, we used a tissue-mimicking phantom with 1.3 nM SERS NPs inside Tygon tubing. This high concentration may be achieved by magnetic trapping of tens to hundreds of cells labeled with hundreds to thousands of SERS NPs. Figure 5 shows the photograph setup used to measure maximum penetration depth. The maximum depth of penetration was 2 mm in a solution of intralipid with the reduced scattering coefficient measured as . Therefore, the maximum depth of penetration in tissue with an of is . 3.2.CrosstalkTo investigate multiplexing capabilities, we quantified crosstalk from varying types of potential interfering SERS NPs. Using multiple concentrations from 10 pM to 1.3 nM of S440 SERS NPs, the estimated concentration of S420 SERS NPs was below the noise of the system when no S420 SERS NPs were present. Likewise, using multiple concentrations from 10 pM to 1.3 nM of S420 SERS NPs, the estimated concentration of S440 SERS NPs was below the noise of the system when no S440 SERS NPs were present. Further, using multiple concentrations of mixed S440 and S421 SERS NPs with the concentration ratio of , the data present highly linear relationships between estimated SERS NP concentrations and actual SERS NP concentrations with (Fig. 6) down to of SERS NP S440. Other groups have performed additional investigations to demonstrate that 10 or more variants of SERS NPs can be demixed with low crosstalk.15 3.3.Nonspecific BindingTo quantify the specificity of binding of folate-conjugated SERS NPs to folate receptors, we used a commercial Raman microscope (Nicolet Almega XR Micro and Macro Raman Analysis System, Thermo Fisher Scientific Inc., Waltham). This permitted us to focus on single cells or groups of cells mounted on a slide after three washing steps, while avoiding possible SERS NPs in the surrounding media. Six samples were prepared by using HeLa (high folate receptor expression) and ZR-75-1 (low folate receptor expression) cell lines. HeLa cells and ZR-75-1 cells were incubated for 4 h, respectively, with: (a) medium only, (b) nonfunctionalized SERS NPs in medium, and (c) folate-conjugated SERS NPs in medium. The SERS spectrum detected from the six samples are shown in Fig. 7. HeLa cells incubated with folate-conjugated SERS NPs generated much higher SERS signals than the other five samples, which proves that folate receptor-positive cells accumulated more folate-conjugated nanoparticles. The detected SERS signal from HeLa cells was 12 times higher than the SERS signal from ZR-75-1 cells. This nonspecific binding ratio may vary depending on the cell type, and it is expected that specificity due to antibody or peptide targeting could be significantly higher. SERS signal from red blood cells mixed with equivalent levels of folate-targeted SERS NPs was not measurable. In addition, fluorescence and Raman microscopy validate that both MNPs and SERS NPs were bound to HeLa cells even after washing steps. Fig. 7SERS signal due to 780-nm excitation wavelength from HeLa cells incubated with medium only, HeLa cells incubated with SERS NPs in medium, HeLa cells incubated with folate targeted SERS NPs in medium, ZR-75-1 cells incubated with medium only, ZR-75-1 cells incubated with SERS NPs in medium, and ZR-75-1 cells incubated with folate targeted SERS NPs in medium, respectively. 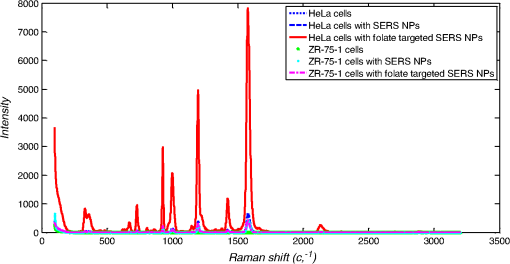 3.4.Magnetic Trapping at 1 cm / s Mean Flow VelocityHeLa cells () were incubated with both folate-conjugated SERS NPs and MNPs (magnetic-fluorescent bead) for 4 h, with fluorescence images demonstrating MNP targeting on HeLa cells. Possible shorter incubation times and in vivo targeting need to be explored in the future. The mean flow velocity (defined as volumetric flow rate divided by cross-sectional area) of HeLa cells mixed with 10 mL PBS inside the 1-mm-inner-diameter Tygon tubing was set as . The detected SERS spectrum changing with the magnetic trapping time is shown in Fig. 8. SERS signal intensity was determined by the magnetically trapped HeLa cells (incubated with both SERS NPs and MNPs) in the optical focal volume, hence the SERS NPs concentration at the focus. For comparison, SERS signals from samples of HeLa cells incubated with folate-conjugated SERS NPs only, and HeLa cells incubated with folate-conjugated MNPs only were detected, as shown in Fig. 9. It is clear that only HeLa cells incubated with both folate-conjugated SERS NPs and MNPs exhibited increasing SERS signals with magnetic trapping time. No such increase was observed in the absence of a magnetic field gradient. Fig. 8SERS spectrum from HeLa cells targeted with both SERS NPs and MNPs at different magnetic trapping times for flow velocity inside 1-mm-diameter Tygon tubing. 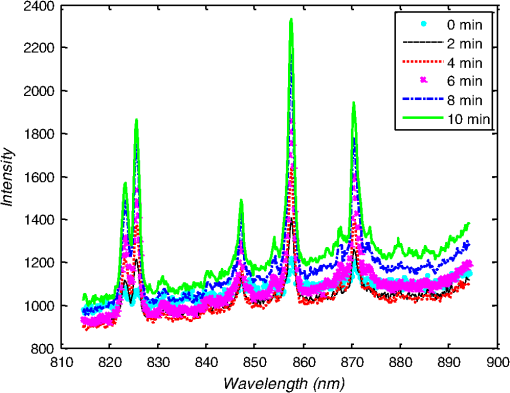 Fig. 9Estimated SERS NP concentration at focal zone detected at different magnetic trapping times for flow velocity inside 1-mm-diameter Tygon tubing. Three types of cell preparations were made: HeLa cells incubated with SERS NPs only, HeLa cells incubated with MNPs only, and HeLa cells incubated with both SERS NPs and MNPs, respectively. 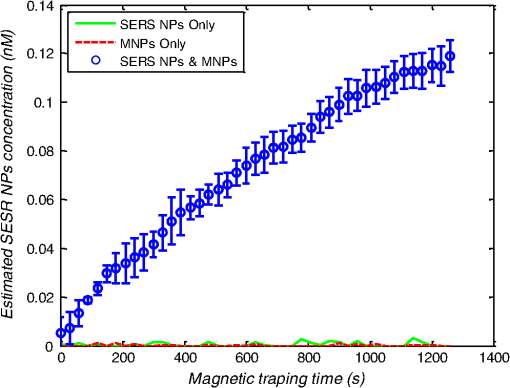 Magnetic trapping and SERS detection were performed using different concentrations of labeled HeLa cells as shown in Fig. 10. After incubation with both folate-conjugated SERS NPs and MNPs, serial dilutions corresponding to , , , , and of HeLa cells were resuspended in 10 mL PBS. These serial diluted samples were flowed through 1-mm-diameter Tygon tubing at mean flow velocity. The estimated SERS NP concentrations in the focal zone increased with magnetic trapping time as shown in Fig. 10, while Fig. 11 presents the details for three low cell concentration samples. Discrete trapping events were observed in the SERS signal for low cell concentration samples, which we believe were due to trapped cells rather than free NPs or NPs washed off the cells, since only HeLa cells dual-labeled with both folate-conjugated SERS NPs and MNPs were detectable as shown in Fig. 9. The discrete increases of estimated SERS NP concentration were at least 10 pM, which corresponds to SERS NPs trapped in the focal zone. Because there may be hundreds or more SERS NPs per cell, the discrete increases in SERS signals are most likely due to single cell trapping or trapping of cell clusters. Variability in the step increases could be due to NP loading variance or cell clustering; however, if cells are loaded with SERS NPs, we are sensitive to single cell trapping. The result also shows that we are able to detect in a total volume of 10 mL within 7 min. Detection of clinically relevant levels of CTCs ( to ) can be achievable if longer magnetic trapping times or larger volumes are used. Figure 12 demonstrated the relationship between estimated SERS NP concentrations in focal zone with cell concentrations, using the same data for Fig. 10, at magnetic trapping time of 900 and 1200 s, respectively. The data present a highly linear relationship between estimated SERS NP concentrations and cell concentrations in PBS solution with a coefficient of determination . For in vivo applications, quantitation of nanoparticle concentrations or CTCs may be more challenging than present ex vivo experiments. 3.5.Magnetic Trapping at Different Flow VelocitiesWe investigated magnetic trapping at different flow velocities ranging from 0.2 to as shown in Fig. 13. These flow velocities are representative of physiological flow velocities in peripheral vessels where future magnetic trapping experiments could be conducted in vivo. It can be seen that the magnetic trapping speed increased with the increasing flowing velocity. Therefore, our system demonstrated its ability to trap flowing cells within a large flow velocity range. 3.6.Detection of Discrete Trapping Events in BloodApproximately 100,000 dual-nanoparticle-labeled HeLa cells were mixed in 10 mL of rat blood. The mixture was flowed through 1-mm-inner-diameter Tygon tubing at a flow velocity of with both incident light and the cone magnet directed at the top of the tubing. Discrete trapping events were observed in the SERS signal shown in Fig. 14. 4.DiscussionTo our knowledge, this is the first report of using both SERS NPs and MNPs to achieve magnetic enrichment and trapping of cancer cells mixed with circulating PBS or blood. The present work offers significant promise for future SERS-based multiplex detection of CTCs and could have both ex vivo and in vivo applications. In vivo clinical applications could involve a subject wearing a magnet for an extended period of time to ensure trapping of adequate numbers of CTCs. For ex vivo applications, CTC detection in blood samples might be considered similar to present work. Specificity can be further improved by the use of antibodies or peptides and the targeting of multiple ligands. SERS signals from HeLa cells targeted with only SERS NPs or only MNPs were well below the sensitivity of our system, while the SERS signals from HeLa cells targeted with both SERS NPs and MNPs increased with the magnetic trapping time. The unity-SNR sensitivity was . The minimum number of SERS NPs detected within our optical focus is estimated to be only , lower than that reported in Ref. 20, which is 600. Nie and Emory21 reported the detection of single SERS NPs. Our data suggest we can detect trapping of single cells labeled with hundreds of NPs. Indeed, brighter SERS NPs and enhanced-sensitivity detection may offer future improvements. Currently, for simplicity, we use only a few spatial channels ( vertical pixels) of our imaging spectrometer system. The detection limit can potentially be further improved by using a fiber bundle to fully utilize all possible spatial channels on our EMCCD and by improving matching of NAs between our spectrometer and detection optics. Additionally, the laser power used in our experiment was about 54 mW measured after the objective lens, slightly less than other groups.15 In the future, improved signal-to-noise or deeper penetration might be achieved by increasing the laser power. Using the long dimension of our EMCCD for spectral discrimination may facilitate a wider spectral range to improve signal-to-noise and enhance multiplexing capability. The overexpression of folate receptor on HeLa cells was reported as .22 In our experiments, the ratio of SERS NPs to HeLa cells was approximately 20,000 to 1. Since the minimum number of SERS NPs detected within our optical focus is estimated to be only , only 1 cell needs to be trapped in order to be detectable using our system if there were 200 SERS NPs targeted on each HeLa cell. This also helps to explain the discrete trapping events observed in Figs. 11 and 14. The discrete increases of estimated SERS NP concentration were at least 10 pM, which corresponds to more SERS NPs trapped in the focal zone. The discrete trapping events indicate one cell or several cells simultaneously trapped by the magnet. Our present study is limited to using tumor cells prelabeled with nanoparticles. Future work should optimize protocols for targeting CTCs in whole blood, both ex vivo and in vivo. Serial dilution experiments indicate it should be possible to detect low clinically relevant concentrations of CTCs using prolonged trapping times. Since folate receptor is overexpressed in many cancer cells, MNPs decorated with folate might target several kinds of tumor cells. Therefore, decorating several flavors of SERS NPs with different antibodies can be potentially used for targeting different types of tumor cells to achieve multiplex detection. Targeting different receptors with multiple SERS NPs may additionally enhance specificity of a single CTC cell type. Immune stealth should also be investigated. For some applications including ex vivo procedures, immune-suppressants could mitigate immune-cell nonspecific uptake. If extraction methods are developed, the enriched population of cells could be further sorted with alternative techniques such as flow cytometry. Although a number of additional research directions appear promising, the present study offers critical feasibility to motivate future work. 5.ConclusionWe have demonstrated magnetic enrichment and detection of CTCs with folate-conjugated MNPs and SERS NPs. This technique could be used in either ex vivo blood samples or in vivo, where a patient could wear a magnet for a sufficient time to detect adequate numbers of CTCs. We hope that present ex vivo experiments may pave the way to create new tools in the detection and, therefore, prediction of high-risk malignancy that requires early aggressive systemic therapy. AcknowledgmentsThe authors gratefully acknowledge funding from a Prostate Cancer Canada Movember Discovery Grant, NSERC (355544-2008, 375340-2009, and STPGP 396444), the Canadian Cancer Society (CCS 2011-700718), the Canada Foundation for Innovation, Leaders Opportunity Fund (18472), Alberta Advanced Education and Technology, Small Equipment Grants Program (URSI09007SEG), Alberta Ingenuity and Alberta Innovates scholarships for graduate students, Canadian Federation of University Women Edmonton Margaret Brine Graduate Scholarships for Women, and SPIE Optics and Photonics Education Scholarship. ReferencesB. WeigeltJ. L. PeterseL. J. van’t Veer,
“Breast cancer metastasis: markers and models,”
Nat. Rev. Cancer, 5
(8), 591
–602
(2005). http://dx.doi.org/10.1038/nrc1670 NRCAC4 1474-175X Google Scholar
R. A. GhosseinS. BhattacharyaJ. Rosai,
“Molecular detection of micrometastases and circulating tumor cells in solid tumors,”
Clin. Cancer Res., 5
(8), 1950
–1960
(1999). CCREF4 1078-0432 Google Scholar
M. C. MillerG. V. DoyleL. W. Terstappen,
“Significance of circulating tumor cells detected by the cellsearch system in patients with metastatic breast colorectal and prostate cancer,”
J. Oncol., 2010 617421
(2010). http://dx.doi.org/10.1155/2010/617421 JOONA2 1687-8469 Google Scholar
R. F. SwabyM. Cristofanilli,
“Circulating tumor cells in breast cancer: a tool whose time has come of age,”
BMC Med., 9 43
(2011). http://dx.doi.org/10.1186/1741-7015-9-43 BMMECZ 1741-7015 Google Scholar
D. C. DanilaM. FleisherH. I. Scher,
“Circulating tumor cells as biomarkers in prostate cancer,”
Clin. Cancer Res., 17
(12), 3903
–3912
(2011). http://dx.doi.org/10.1158/1078-0432.CCR-10-2650 CCREF4 1078-0432 Google Scholar
F. Tanakaet al.,
“Circulating tumor cell as a diagnostic marker in primary lung cancer,”
Clin. Cancer Res., 15
(22), 6980
–6986
(2009). http://dx.doi.org/10.1158/1078-0432.CCR-09-1095 CCREF4 1078-0432 Google Scholar
B. P. NeginS. J. Cohen,
“Circulating tumor cells in colorectal cancer: past, present, and future challenges,”
Curr. Treat. Opt. Oncol., 11
(1–2), 1
–13
(2010). http://dx.doi.org/10.1007/s11864-010-0115-3 1527-2729 Google Scholar
W. J. Allardet al.,
“Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases,”
Clin. Cancer Res., 10
(20), 6897
–6904
(2004). http://dx.doi.org/10.1158/1078-0432.CCR-04-0378 CCREF4 1078-0432 Google Scholar
S. Nagrathet al.,
“Isolation of rare circulating tumor cells in cancer patients by microchip technology,”
Nature, 450 1235
–1239
(2007). http://dx.doi.org/10.1038/nature06385 NATUAS 0028-0836 Google Scholar
K. Kazunoriet al.,
“Microchip-based immunomagnetic detection of circulating tumor cells,”
Lab Chip, 11
(20), 3449
–3457
(2011). http://dx.doi.org/10.1039/c1lc20270g LCAHAM 1473-0197 Google Scholar
S. Zhenget al.,
“Membrane microfilter device for selective capture, electrolysis and genomic analysis of human circulating tumor cells,”
J. Chromatogr., 1162
(2), 154
–161
(2007). http://dx.doi.org/10.1016/j.chroma.2007.05.064 JOCRAM 0021-9673 Google Scholar
E. I. Galanzhaet al.,
“In vivo magnetic enrichment and multiplex photoacoustic detection of circulating tumor cells,”
Nat. Nanotechnol., 4 855
–860
(2009). http://dx.doi.org/10.1038/nnano.2009.333 1748-3387 Google Scholar
C. W. Weiet al.,
“Trapping and dynamic manipulation of polystyrene beads mimicking circulating tumor cells using targeted magnetic/photoacoustic contrast agents,”
J. Biomed. Opt., 17
(10), 101517
(2012). http://dx.doi.org/10.1117/1.JBO.17.10.101517 JBOPFO 1083-3668 Google Scholar
C. M. O’Brienet al.,
“Capture of circulating tumor cells using photoacoustic flowmetry and two phase flow,”
J. Biomed. Opt., 17
(6), 061221
(2012). http://dx.doi.org/10.1117/1.JBO.17.6.061221 JBOPFO 1083-3668 Google Scholar
C. L. Zavaletaet al.,
“Multiplexed imaging of surface enhanced Raman scattering nanotags in living mice using noninvasive Raman spectroscopy,”
Proc. Natl. Acad. Sci. U. S. A., 106
(32), 13511
–13516
(2009). http://dx.doi.org/10.1073/pnas.0813327106 PNASA6 0027-8424 Google Scholar
X. Wanget al.,
“Detection of circulating tumor cells in human peripheral blood using surface-enhanced Raman scattering nanoparticles,”
Cancer Res., 71
(5), 1526
–1532
(2011). http://dx.doi.org/10.1158/0008-5472.CAN-10-3069 CNREA8 0008-5472 Google Scholar
B. Junet al.,
“Multifunctional silver-embedded magnetic nanoparticles as SERS nanoprobes and their applications,”
Small, 6
(1), 119
–125
(2010). http://dx.doi.org/10.1002/smll.v6:1 1613-6829 Google Scholar
P. Quaresmaet al.,
“Star-shaped magnetite@gold nanoparticles for protein magnetic separation and SERS detection,”
RSC Adv., 4
(8), 3659
–3667
(2013). http://dx.doi.org/10.1039/c3ra46762g RSCACL 2046-2069 Google Scholar
P. S. LowA. C. Antony,
“Folate receptor-targeted drugs for cancer and inflammatory disease,”
Adv. Drug Deliv. Rev., 56
(8), 1055
–1231
(2004). http://dx.doi.org/10.1016/j.addr.2004.02.003 ADDREP 0169-409X Google Scholar
S. Kerenet al.,
“Noninvasive molecular imaging of small living subjects using Raman spectroscopy,”
Proc. Natl. Acad. Sci. U. S. A., 105
(15), 5844
–5849
(2008). http://dx.doi.org/10.1073/pnas.0710575105 PNASA6 0027-8424 Google Scholar
S. NieS. R. Emory,
“Probing single molecules and single nanoparticles by surface-enhanced Raman scattering,”
Science, 275
(5303), 1102
–1106
(1997). http://dx.doi.org/10.1126/science.275.5303.1102 SCIEAS 0036-8075 Google Scholar
G. Destitoet al.,
“Folic acid-mediated targeting of cowpea mosaic virus particles to tumor cells,”
Chem. Biol., 14
(10), 1152
–1162
(2007). http://dx.doi.org/10.1016/j.chembiol.2007.08.015 CBOLE2 1074-5521 Google Scholar
Biography |