|
|
1.IntroductionThe advent of two-photon excitation microscopy (TPEM) has revolutionized biological imaging by enabling the detection of dynamic cellular processes in intact tissues deep into the organs of living organisms. When combined with the use of fluorescent indicators, TPEM can be used to follow the variation of intracellular [] within the cells in their physiologic environment (i.e., the tissue). In the heart, intracellular is a key determinant of cardiomyocyte function, which is regulated by a complex of signals shaped in time (from millisecond to second) and space (from subcellular to cell-wide) within the single cell. In addition, fast and chaotic signals occurring during arrhythmias involving, by definition, multiple cells in the tissue add another layer of complexity as cell–cell interactions take part in the arrhythmia pathophysiology. The ideal method to visualize intracellular fluctuations in the intact myocardium would thus be endowed with high temporal and spatial resolutions and, preferentially, be able to record the cellular fluorescence from a wide field of view. Up until now, one of the limitations of conventional TPEM is the difficulty in acquiring full frame () images at a frame rate significantly higher than 1 Hz, thus hampering the simultaneous detection of transient dynamic events lasting fractions of a second in a wide sample region.1,2 Different solutions have been used3 to achieve high speed and resolution while retaining the advantages of two-photon excitation (TPE), such as penetration depth and reduced out of focus phototoxicity, with respect to the traditional confocal or fluorescent microscopy.4 Reduced scanned area and/or pixel resolution are the straightforward ideas developed by microscopy companies and are normally used by several research groups to increase the image acquisition rate.5–7 However, all these strategies allow faster acquisition at the expense of the spatial details, up to the most extreme case of a single-line scan that uses the deflector to sweep the laser beam along a single line, thus generating two-dimensional () images. One alternative strategy to achieve full-field imaging at a high rate would exploit faster beam scanners and compensate for the inherent reduction in pixel dwell time by intensifying the incident laser power. However, given that the photodamage of biological specimens excited at two photons increases nonlinearly with the laser power,8 this last approach has very little use in living samples. To reduce photodamage and photobleaching, Konnerth and collaborators demonstrated the possibility to shorten the dwell time without compromising the signal using low-power temporal oversampling. In this technique, a mirrorless scanning two-photon microscope is used and has been applied to image signals in vivo.9,10 Other techniques exploiting acousto-optic deflectors instead of galvanometric mirrors have been applied to imaging11 but do not solve the problem of high resolution in a large field of view. An alternative and promising approach to image thick samples while maintaining large field of view consists of the combination of TPEM and selective plane illumination microscopy. This technique has been used for developmental studies12 and is fully characterized for highly scattering samples.13 To increase the acquisition rate of TPEM, we used a parallel scanning system based on the insertion of a diffractive optical element (DOE)14 along the incident laser path, splitting the beam into a matrix of identical beamlets with a well-defined spatial distribution. By using such an arrangement, the sample area was divided into a number of contiguous subregions, each of which was simultaneously raster scanned by a single beamlet. Parallel acquisition by an array of detectors, with a geometry corresponding to that of the DOE, allowed the spatial resolution of the detection path and the reconstruction of the full-filed sample image. The multiphoton multispot microscopy system described in this work (MCube, developed by light4tech15) was installed at the Department of Biomedical Sciences and was used to test and compare image quality and point spread function (PSF) with those of a conventional TPEM system.16 Subsequently, the MCube was used for imaging experiments in acute heart slices, allowing the simultaneous detection of subcellular dynamics in several different cells in the tissue. The cardiac slices’ model was selected as a thick-tissue testing platform for multiphoton microscopy as it combines the preservation of the multicellular myocardial structure with the quasi-bidimensional morphology of the slice.17 2.Materials and Methods2.1.Conventional Two-Photon Excitation MicroscopeA multiphoton system (Radiance 2100 MP; Bio-Rad Laboratories, UK) was used for the comparison of MCube image quality with respect to an existing TPE microscopy system. The radiance system is equipped with a Tsunami mode-locked, tunable, femto-second–pulsed Ti/sapphire laser (Ti:Sa). The laser output is capable of generating 100-fs pulse trains at a rate of 82 MHz. The microscope (Eclipse E600FN; Nikon) was equipped with a Nikon objective (); a direct detection system (Bio-Rad), fitted with a 500LP DC dichroic mirror and an HQ535/50 emission filter (Chroma Technology Corp.), was used for the detection of fluorescence emission signals. The LaserSharp2000 software package (Bio-Rad) was used for data acquisition. 2.2.Acute Heart Slices Cutting and LoadingAcute ventricular heart slices were cut following the protocol described in Ref. 18. Young mice (postnatal day 7 to 10) were sacrificed and the heart was quickly excised and washed in ice-cold -free Tyrode solution (composition in mM: NaCl 136, KCl 5.4, 0.33, 1, glucose 10, hepes 5; pH 7.40 with NaOH). Atria were excised by a cut on the transversal plane, also creating a basis surface for fixing the ventricular portion of the heart to the slicer support with a commercial cyanoacrylate glue. Hearts from young mice were embedded in 4% low-melting agarose (SIGMA-Aldrich) in Tyrode. Transversal slices ( thickness) were cut in ice-cold -free Tyrode solution with a vibratome (Leica GmBH). After cutting, slices were transferred to room temperature 1-m M Tyrode for some minutes and then they were maintained in the recovery medium (DMEM/F12 supplemented with 20% Knock-out Replacement Serum, Invitrogen) in a humidified atmosphere containing 95% and 5% at 37°C for at least 1 h before loading.19 After recovery, slices were loaded with the -sensitive indicator Fluo-4AM (, Invitrogen) for 40 min at 37°C in 1-mM Tyrode solution plus 20% Pluronic F-127 (Invitrogen) and sulfinpyrazone (SIGMA-Aldrich). 2.3.Imaging and StimulationViable heart slices loaded with Fluo-4AM were transferred on a homemade perfusion chamber on the stage of the two-photon microscope and perfused with a 1 mM , oxygenated Tyrode solution. To prevent curling and movement, slices were held down by a homemade platinum holder. Slices were field stimulated by applying directly under the microscope a voltage difference at the slice sides (); slices were thus electrically paced by applying brief voltage square waveforms (5 ms), with the fixed frequency of 1, 2, or 5 Hz. Changes in concentration were observed as variations in the green emission of the sample upon excitation. 3.Results3.1.Instrument DesignMultispot multiphoton microscopy (MMM) allows the increase of the image acquisition speed several fold when compared with standard laser scanning microscopes. Analogous systems based on a microlens array have already been used to monitor dynamics in isolated cardiac cells7 but, to the best of our knowledge, MMM has never been employed to image cellular in whole hearts or ex vivo myocardial preparations. To obtain an MMM system suitable for high-speed imaging at the cellular level in intact organs, we sought to obtain homogeneous light distribution over the sample, while at the same time, maximize light detection from the whole field of view of the objective. The system design is thus based on a DOE in the excitation light path, a descanned configuration in the emission path, and an array of photomultiplier tubes (PMTs) [Fig. 1(a)]. A Ti:Sa pulsed infrared (IR) laser (, 80-MHz pulses, Chameleon Ultra II, Coherent, UK) providing up to 3.6 W of optical power at 785-nm excitation wavelength was used as the excitation light source. Fig. 1(a) Scheme of the multispot multiphoton microscope (MMM). (b) Essential components of the MMM. (c) Image and intensity profile of a subresolution bead along the radial axis. 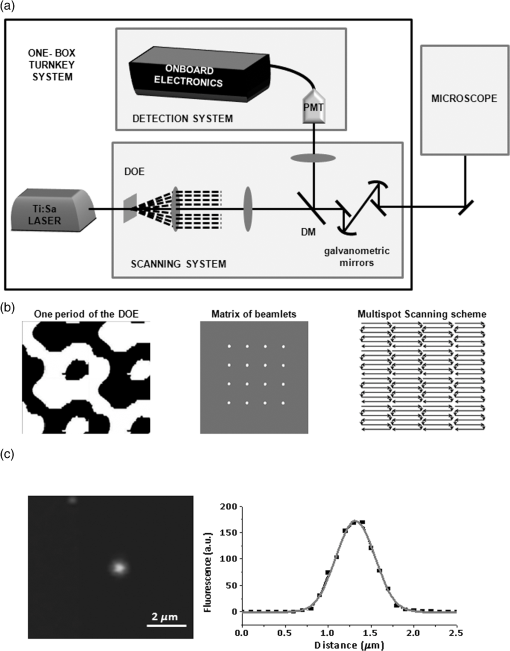 The DOE was inserted along the incident light path to obtain patterned illumination of the sample, obtained by splitting the laser beam into 16 parallel beamlets generating a matrix of 4 by 4 spots regularly spaced in a squared array. Such a configuration, in the preliminary tests, offered the best compromise between the imaging rate and signal-to-noise ratio. Laser power was evenly distributed throughout the spots [within a few percent due to the DOE construction, Fig. 1(b)], thus providing uniform illumination of the entire scanned area.14 With the laser power at the source of 3.6 W, the power per beamlet measured after the DOE resulted in about 45 mW. The 16 parallel beamlet patterns were deflected along the and axes by galvanometric mirrors, while movement of the excitation plane along the -axis was achieved by shifting the objective with a piezoelectric motor. Detection of the emitted light is performed with a multianode PMT, (H7546A-20SEL from Hamamatsu), with each detecting element corresponding to a single descanned beamlet. Each PMT has its own electronic board, performing signal amplification, A/D conversion, local storage on a 2 MB onboard memory, and parallel communication with a custom-developed control software for data acquisition and storage on the PC hard drive. This configuration minimizes the optical cross-talk between different PMT elements20 that may introduce undesired degradation of the image quality, particularly when acquiring fluorescence from highly scattering samples such as muscular tissue. The essential components of the MMM are shown in Fig. 1(a). To allow experiments with living tissue samples, the MMM was adapted to an upright fluorescence microscope (DM LFSA, Leica Microsystems GmBH, Germany), equipped with a water immersion objective (Nikon, ). 3.2.Optical PerformanceTo evaluate the optical performance of the MMM system, we determined the PSF by collecting the three-dimensional (3-D) image profile of a subresolution fluorescent bead, as shown in Fig. 1(c). From the radial intensity profile in Fig. 1(c), we obtained a radial resolution of , a value in line with the expected theoretical resolution of 455 nm calculated for a two-photon system at the excitation wavelength of 785 nm. Imaging of fluorescent beads embedded in agarose has previously been used to measure the PSF of microscopy systems as shown in Refs. 7 and 20. Although such a configuration is adequate for evaluating the pure optical performance of the systems, the sample is almost transparent and isotropic, and thus is not representative of the typical experimental condition. The myocardium is densely occupied by sarcomeres, formed by alternating segments of thin and thick protein filaments arranged in a highly scattering grid, inevitably reducing the performance of an optical system. To test the optical performance of the MMM system in deep tissue imaging in the heart, we evaluated the experimental PSF using a custom-generated heart phantom sample. The phantom was made by injecting fluorescent beads (Invitrogen) at different positions and depths in a fixed heart slice. We measured the fluorescence profile of a bead at a depth in the tissue and estimated the system PSF in deep tissue imaging. The ePSF obtained is shown in Fig. 2(a) and the characteristics of fluorescence profiles are reported in Table 1. Fig. 2(a) Intensity profile of a fluorescent bead at depth in myocardial tissue. Scale bars: . Radial and axial views are reported as well as relative intensity profiles. (b) The same image acquired in single-line modality and multispot modality.  Table 1Full width at half maximum (FWHM) of the measured point spread functions (PSFs) for the single-beam BIORAD MP2100 and multispot MCube systems; data refer to a fluorescent bead located 150 μm in depth in the phantom sample.
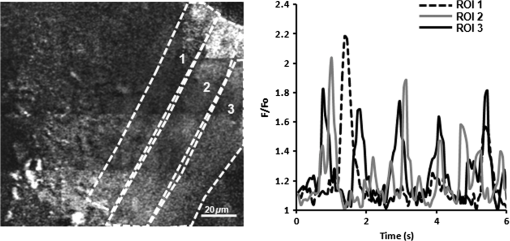
The results obtained with the MMM were compared with that of a commercial standard, the BIORAD MP2100 multiphoton microscope powered by a Ti:Sa Spectra Physics Millennia pulsed IR laser. We obtained, in both cases, a PSF for TPE microscopy comparable with that reported in other papers,7 with a better resolution for the MMM in both the axial and radial directions. The improvement in the optical resolution of the MMM can be dependent, at least in part, on the spatial filtering of the DOE apparatus. To determine the effect of the DOE insertion, we next compared the quality of the images acquired in either single-line mode or multispot mode of the same field of view with a comparable laser power. In a single-line mode, the laser power was thus attenuated down to 45 mW on the sample plane. As shown in Fig. 2(b), the image resolution and S/N are comparable, demonstrating that the DOE insertion does not cause image deterioration. Notably, the acquisition with the DOE inserted is 16 times faster than the single-line modality. In the inset, a magnification is shown. 3.3.Multispot Multiphoton Microscopy Imaging of Calcium Dynamics in Thick Myocardial SlicesThe system was tested on viable acute cardiac slices from mouse heart, by performing imaging using the loaded fluorescent calcium indicator Fluo-4AM. Acute myocardial slices are an experimental system well representing the intact myocardium, allowing the use of state-of-the art imaging methods to explore signals from the subcellular to the multicellular range. We used the MMM system to perform two different sets of experiments. In the first, imaging was performed at 16-Hz rate during field stimulation in a wide area of the myocardial slice, typically including six to eight cardiomyocytes. Electrical pacing by field stimulation resulted in a synchronous increase in intracellular in the cardiomyocytes in the imaging field, as reported in a typical experiment shown in Fig. 3(a). At the acquisition rate used in the experiments, it was possible to fit the transients’ decay with a single exponential function obtaining a typical decay time in the range of 100 to 150 ms (), in line with data in the literature.21 In the subsequent set of experiments, MMM was used to investigate dynamics with a subcellular resolution in a networked group of cells in the myocardial slices. Fig. 3(a) Intensity profiles of three neighboring cells in the heart slice. Cell borders are highlighted by dotted squares and the fluorescence profiles are reported. It can be seen that cell 2 crosses two subframes without affecting temporal fluorescence profiles. (b) Simultaneous onset or macrosparks in two of the three regions of interest (ROI) considered with the respective intensity profiles. (c) A travelling wave recorded in the same cell and intensity profiles of the ROIs considered. 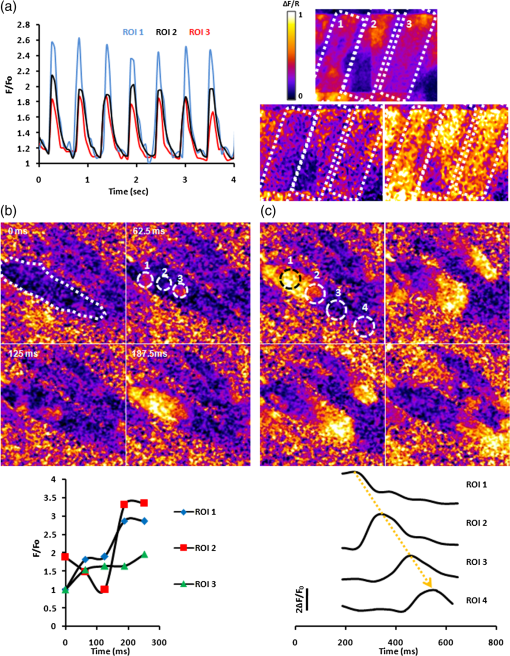 The spatial resolution of MMM at a 16 Hz frame rate was assessed by imaging subcellular dynamics in a wide region of the slice. The elementary signals regulating the excitation–contraction coupling in heart cells are represented by “sparks,”22 i.e., transient and localized releases of from the sarcoplasmic reticulum (SR) through the ryanodine receptor channels. A pace-stop protocol was used to increase the frequency of release events. Intracellular and local release events, both in the form of secondary “macrosparks” [Fig. 3(b)] of approximately 4 to 6 μm in diameter, and self-sustained waves traveling through the cell [Fig. 3(c)] could be imaged simultaneously in numerous cardiomyocytes in the slice. Such cell-wide waves frequently originated from the same release hotspot and, in some cases, were underlined by organized intracellular rotors (see Fig 4). Moreover, it was possible to appreciate that the neighboring cells displayed waves in a pattern of interdependency (Fig. 5). Fig. 4Travelling waves and rotors in the same region of Fig. 3(b) (MOV, 7.93 MB). [URL: http://dx.doi.org/10.1117/1.JBO.20.5.051016.1.] 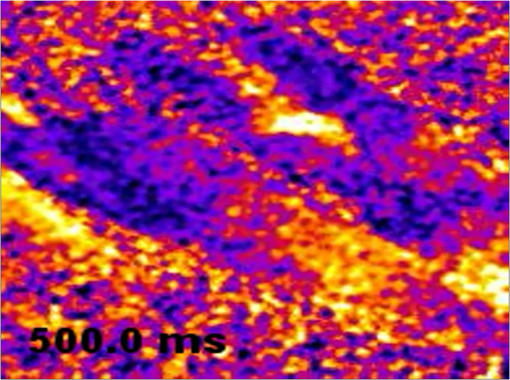 4.DiscussionThe simultaneous analysis of intracellular dynamics in several neighboring cells in their physiologic environment, using fluorescence-based methods, poses a number of technical challenges:
In standard laser scanning confocal microscopes and TPEM, a single collimated laser beam is scanned on the sample in a raster pattern. The maximal frame rate that can be achieved in such systems is essentially determined by two parameters: (1) the speed of the beam deflectors, typically galvanometric mirrors. (2) the pixel dwell time, determined by the shortest time necessary to achieve sufficient excitation of the sample. The strategy chosen to increase acquisition speed was to split the laser beam into a grid of beamlets focused in parallel portions of the sample via a DOE positioned along the laser pathway. If compared with other multispot microscopes in which, for example, the beam is split using a microlens array,7 the DOE insertion guarantees a homogeneous distribution of the light intensity over the sample, thus reducing the need for off-line processing of the images.14 This detection scheme has the advantage with respect to a CCD that optical cross-talk between the different fluorescence channels is strongly reduced.23 This is essential when deep imaging is performed and cross-talk artifacts may completely destroy the image quality.20 For this reason, in order to reconstruct a single output image with a 16 PMT microscope, it is crucial that the PMTs are equalized to the output intensity for identical input brightness. The PMTs of the detection system have been optimized for measurements of calcium dynamics in thick tissue samples and then used for the first time for imaging of acute cardiac slices. The choice of a detection scheme exploiting parallel detection performed by an array of PMTs was essential to allow deep imaging, avoiding cross-talk artifacts. However, each PMT is an independent component having different gain efficacies. To compensate for the intrinsic differences of individual PMTs, all the tubes can be equalized by independently adjusting gain and offset. The signal level can be equalized within 1%, thus compensating any source of nonuniformity in the excitation/detection path. Notwithstanding these adjustments, perfect equalization of the PMTs could not be obtained by us and others, likely due to the inherent differences in the noise level characteristic of each PMT. Here, we showed that the multispot scanning system applied to a multiphoton microscope greatly improves its temporal resolution without worsening the spatial resolution. First, we characterized microscope resolution by calculating radial and axial resolutions in a thick tissue phantom. This allowed us to verify that the laser could penetrate deep enough inside the cardiac tissue without losing the contrast; this is important for our study because we need to enter inside the sample for at least (around two to three cells) to overcome the surface directly damaged by the cutting procedure. It has to be considered that all the -imaging experiments were performed below the slice surface, where we also measured the PSF, in order to overcome the cell layers suffering from secondary dysfunctions. Once assessed, the microscope resolution of the experimental model of acute ventricular slices of murine heart has been implemented and optimized for imaging measurements in wide myocardial areas. The feasibility of ventricular heart slices has already been reported;18,19 nonetheless, this model has been previously used to monitor electrophysiological parameters and as a useful model for pharmacological drug testing,24 but much less for imaging. Acute myocardial slices are a good model for fluorescence imaging because they associate the ease of the bidimensional exposition of cells under the objective with the intrinsic complex network of a 3-D tissue. A limitation in the use of murine heart slices for prolonged physiological measurements may relate to oxygen diffusion in the tissue. To reduce the impact of such a problem, we have perfused the tissue slice on both sides and used short stimulation protocols at room temperature, as suggested by Barclay with respect to a similar cardiac tissue preparation.25 In this work, we showed that an MMM can be exploited not only for morphological studies in thick tissues as previously reported,23 but also to study secondary messengers in tissue preparations or whole organs. As we showed, the possibility to acquire full images at 16 Hz without worsening the spatial resolution opens new possibilities for studies of dynamics in intact tissues including the heart. Specifically, in cardiomyocytes physiology, releases of from the SR, sparks, are fundamental in the excitation–contraction coupling.22 Moreover, abnormal releases from the SR have been associated with different channelopathies,26,27 and the possibility of studying subcellular events in the intact tissue is now possible with a full-frame approach allowing the detection of macrosparks, waves, and rotors. The use of TPE in cardiovascular research is increasing, and the need of methodological improvement is arising.21,28 The properties of MMM will enable the investigation of fast occurring -dependent signals in living cells within intact tissues, thus expanding the methodological toolkit for fluorescence imaging which, in the last 20 years, has become an experimental technique increasingly used in biomedical research.29 AcknowledgmentsThis work was supported by funding from the European Community Seventh Framework Program FP7/2007-2013 under Grant Agreement HEALTH-F2-2009-241526, EUTrigTreat (to M.M. and F.P.). Disclosures: None declared. ReferencesW. DenkJ. H. StricklerW. W. Webb,
“Two-photon laser scanning fluorescence microscopy,”
Science, 248 73
–76
(1990). http://dx.doi.org/10.1126/science.2321027 SCIEAS 0036-8075 Google Scholar
W. R. ZipfelR. M. WilliamW. Webb,
“Nonlinear magic: multiphoton microscopy in the biosciences,”
Nat. Biotechnol., 21
(11), 1369
–1377
(2003). http://dx.doi.org/10.1038/nbt899 NABIF9 1087-0156 Google Scholar
F. HelmchenW. Denk,
“Deep tissue two-photon microscopy,”
Nat. Methods, 2
(12), 932
–939
(2005). http://dx.doi.org/10.1038/nmeth818 1548-7091 Google Scholar
E. HooverJ. Squier,
“Advances in multiphoton microscopy technology,”
Nat. Photonics, 7
(2), 93
–101
(2013). http://dx.doi.org/10.1038/nphoton.2012.361 1749-4885 Google Scholar
T. Nielsenet al.,
“High efficiency beam splitter for multifocal multiphoton microscopy,”
J. Microsc., 201
(3), 368
–376
(2001). http://dx.doi.org/10.1046/j.1365-2818.2001.00852.x JMICAR 0022-2720 Google Scholar
K. SvobodaD. W. TankW. Denk,
“Direct measurement of coupling between dendritic spines and shafts,”
Science, 272
(5262), 716
–719
(1996). http://dx.doi.org/10.1126/science.272.5262.716 SCIEAS 0036-8075 Google Scholar
K. Bahlmannet al.,
“Multifocal multiphoton microscopy (MMM) at a frame rate beyond 600 Hz,”
Opt. Express, 15
(17), 10991
–10998
(2007). http://dx.doi.org/10.1364/OE.15.010991 OPEXFF 1094-4087 Google Scholar
A. HoptE. Neher,
“Highly nonlinear photodamage in two-photon fluorescence microscopy,”
Biophys. J., 80
(4), 2029
–2036
(2001). http://dx.doi.org/10.1016/S0006-3495(01)76173-5 BIOJAU 0006-3495 Google Scholar
X. Chenet al.,
“Functional mapping of single spines in cortical neurons in vivo,”
Nature, 475
(7357), 501
–505
(2011). http://dx.doi.org/10.1038/nature10193 NATUAS 0028-0836 Google Scholar
X. Chenet al.,
“LOTOS-based two-photon calcium imaging of dendritic spines in vivo,”
Nat. Protoc., 7
(10), 1818
–1829
(2012). http://dx.doi.org/10.1038/nprot.2012.106 NPARDW 1754-2189 Google Scholar
C. Crociniet al.,
“ Defects in T-tubular electrical activity underlie local alterations of calcium release in heart failure,”
Proc. Natl. Acad. Sci. U. S. A., 111
(42), 15196
–15201
(2014). http://dx.doi.org/10.1073/pnas.1411557111 PNASA6 0027-8424 Google Scholar
T. V. Trounget al.,
“Deep and fast live imaging with two-photon scanned light sheet microscopy,”
Nat. Methods, 8 757
–760
(2011). http://dx.doi.org/10.1038/nmeth.1652 1548-7091 Google Scholar
Z. Lavagninoet al.,
“Two-photon excitation selective plane illumination microscopy (2PE-SPIM) of highly scattering samples: characterization and application,”
Opt. Express, 21
(5), 5998
–6008
(2013). http://dx.doi.org/10.1364/OE.21.005998 OPEXFF 1094-4087 Google Scholar
L. Sacconiet al.,
“Multiphoton multifocal microscopy exploiting a diffractive optical element,”
Opt. Lett., 28
(20), 1918
–1920
(2003). http://dx.doi.org/10.1364/OL.28.001918 OPLEDP 0146-9592 Google Scholar
A. Diasproet al.,
“Three-dimensional optical behaviour of a confocal microscope with single illumination and detection pinhole through imaging of subresolution beads,”
Microsc. Res. Tech., 45 130
–131
(1999). http://dx.doi.org/10.1002/(SICI)1097-0029(19990415)45:2<130::AID-JEMT7>3.0.CO;2-V MRTEEO 1059-910X Google Scholar
T. P. De Boeret al.,
“Myocardial tissue slices: organotypic pseudo-2D models for cardiac research and development,”
Future Cardiol., 5
(5), 425
–430
(2009). http://dx.doi.org/10.2217/fca.09.32 FCUAAZ 1479-6678 Google Scholar
M. Halbachet al.,
“Ventricular slices of adult mouse hearts—a new multicellular in vitro model for electrophysiological studies,”
Cell. Physiol. Biochem., 18 01
–08
(2006). http://dx.doi.org/10.1159/000095132 CEPBEW 1015-8987 Google Scholar
W. Habeleret al.,
“An in vitro beating model for long-term assessment of experimental therapeutics,”
Cardiovasc. Res., 81 253
–259
(2008). http://dx.doi.org/10.1093/cvr/cvn299 CVREAU 0008-6363 Google Scholar
R. Niesneret al.,
“The power of single and multibeam two-photon microscopy for high-resolution and high-speed deep tissue and intravital imaging,”
Biophys. J., 93
(7), 2519
–2529
(2007). http://dx.doi.org/10.1529/biophysj.106.102459 BIOJAU 0006-3495 Google Scholar
M. Rubartetal.,
“Two-photon molecular excitation imaging of Ca2+ transients in Langendorff-perfused mouse hearts,”
AJP Cell Physiol., 284 1654
–1668
(2003). http://dx.doi.org/10.1152/ajpcell.00469.2002 0363-6143 Google Scholar
H. ChengW. J. LedererM.B. Cannell,
“Calcium sparks: elementary events underlying excitation-contraction coupling in heart muscle,”
Science, 262
(5134), 740
–744
(1993). http://dx.doi.org/10.1126/science.8235594 SCIEAS 0036-8075 Google Scholar
K.H. Kimet al.,
“Multifocal multiphoton microscopy based on multianode photomultipliers tubes,”
Opt. Express, 15
(18), 11658
–11678
(2007). http://dx.doi.org/10.1364/OE.15.011658 OPEXFF 1094-4087 Google Scholar
A. Busseket al.,
“Tissue slices from adult mammalian hearts as a model for pharmacological drug testing,”
Cell. Physiol. Biochem., 24 527
–536
(2009). http://dx.doi.org/10.1159/000257528 CEPBEW 1015-8987 Google Scholar
C.J. Barclay,
“Modelling diffusive O(2) supply to isolated preparations of mammalian skeletal and cardiac muscle,”
J. Muscle Res. Cell Motil., 26 225
–235
(2005). http://dx.doi.org/10.1007/s10974-005-9013-x JMRMD3 0142-4319 Google Scholar
S. R. Houser,
“ signaling domains responsible for cardiac hypertrophy and arrhythmias,”
Circ. Res., 104
(4), 413
–415
(2009). http://dx.doi.org/10.1161/CIRCRESAHA.109.193821 CIRUAL 0009-7330 Google Scholar
S. E. Lehnartet al.,
“Leaky release channel/ryanodine receptor 2 causes seizures and sudden cardiac death in mice,”
J. Clin. Invest., 118
(6), 2230
–2245
(2008). http://dx.doi.org/10.1172/JCI35346 JCINAO 0021-9738 Google Scholar
J. A. ScherschelM. Rubart,
“Cardiovascular imaging using two-photon microscopy,”
Microsc. Microanal., 14
(6), 492
–506
(2008). http://dx.doi.org/10.1017/S1431927608080835 MIMIF7 1431-9276 Google Scholar
R. Rudolfet al.,
“Looking forward to seeing calcium,”
Nat. Rev. Mol. Cell Biol., 4 579
–586
(2003). http://dx.doi.org/10.1038/nrm1153 NRMCBP 1471-0072 Google Scholar
BiographyGiulia Borile received her MSc degree in physics at the University of Padova, where she is completing her PhD in biosciences and biotechnologies, working in the laboratory of Dr. Marco Mongillo at the Venetian Institute of Molecular Medicine. This manuscript is part of her PhD project that involves the use of biophysical approaches to study cardiac arrhythmogenesis with a particular interest in microscopy and Ca2+ imaging. |